
Model of Syntactic Compatibility in Workflows for Electrophysiology
Jan Štěbeták and Roman Mouček
Department of Computer Science and Engineering, University of West Bohemia, Univerzitní 8, Pilsen, Czech Republic
Keywords: Neuroinformatics, Electroencephalography, Event-Related Potentials, Workflows, Analytic Methods,
Syntactic Compatibility, Workflow Steps.
Abstract: Large amounts of EEG/ERP (electroencephalography, event-related potential) data are produced by
scientific laboratories. For complex analysis, data are processed by a set of methods sequentially or in
parallel. These processes are known as workflows. However, various input/output formats of used methods
involve difficulties while putting methods in a pipe. Simple syntactic rules comparing formats of
input/output are already used by workflow engines. In electrophysiology, it is necessary to extend these
rules due to variety of methods. Therefore, extension of syntactic rules between subsequent methods in a
workflow is presented in this paper. The proposed solution allows creating more complex workflows in the
domain of electrophysiology.
1 INTRODUCTION
Our research group specializes in the research of
brain activity; especially attention of drivers is
investigated. We widely use the methods of
electroencephalography (EEG) and event related
potentials (ERP). EEG/ERP experiments usually
take long time and produce a lot of data. Since we
need to analyze experimental data, analytic methods
that we widely use are presented.
For complex analysis, scientists often must
combine multiple processing steps into larger
“analysis pipelines” that can involve a number of
custom algorithms, specialized tools, local and
remote databases, and web services. These “analysis
pipelines” are known as workflows (Littauer, et al.
2012).
In sequential workflows, a result of a previous
method is transferred to a next method. Since putting
methods into workflows is dependent on formats of
input/output of the used methods, the syntactic rules
have to be defined.
In this paper we first briefly describe available
workflows engines and existing ways of ensuring
syntactic compatibility. The next section presents
principles of analytic methods and creating
workflows which are suitable for the electro-
physiology domain. Section 5 describes proposed
extension of ensuring syntactic compatibility
between subsequent methods. A simple comparison
of input/output formats is commonly used in many
workflow engines. However, for complex sequential
workflows in the electrophysiology domain, it is
necessary to use the methods that are incompatible
using a simple syntactic rule. Therefore, we
extended rules that ensure syntactic compatibility. It
consists in defining more formats of input/output
parameters of a method or using a subset of a result
as an input to a next method.
2 STATE OF THE ART
This section briefly describes available workflow
engines and existing ways of ensuring syntactic
compatibility.
2.1 Workflow Engines
The CARMEN project (CARMEN, 2013) has
currently addressed requirements of scientists and
developed a workflow generation and execution
system within the platform. The CARMEN
Workflow Tool is Java-based and designed to make
use of CARMEN Services. The workflow tool
supports both data and control flow, and allows
parallel execution of services. The complete
workflow tool consists of a graphical design tool, a
workflow engine, and access to a library of
CARMEN services and common workflow tasks.
442
Štebeták J. and Moucek R..
Model of Syntactic Compatibility in Workflows for Electrophysiology.
DOI: 10.5220/0004909304420446
In Proceedings of the International Conference on Health Informatics (HEALTHINF-2014), pages 442-446
ISBN: 978-989-758-010-9
Copyright
c
2014 SCITEPRESS (Science and Technology Publications, Lda.)

Taverna (Taverna, 2013) is an open source and
domain-independent Workflow Management System
– a suite of tools used to design and execute
scientific workflows.The Taverna suite is written in
Java and includes the Taverna Engine (used for
enacting workflows) that powers both the Taverna
Workbench (the desktop client application) and the
Taverna Server (which allows remote execution of
workflows). Taverna is also available as
a Command Line Tool for a quick execution of
workflows from a terminal (Taverna, 2013).
e-Science Central is a Cloud based Platform for
Data Analysis. It supports secure storage and
versioning of data, audit and provenance logs and
processing of data using workflows. Workflows are
composed of blocks which can be written in Java, R,
Octave or Javascript (eScience, 2013). Scientists are
able to design workflows using the drag-and-drop
online workflow designer by selecting blocks
(services). The input and output of each block is
typed to prevent incompatible blocks being
connected to each other (Watson, et al. 2010).
2.2 Syntactic Compatibility of
Workflows
The engines described above are designed for
scientific purposes. They provide modelling of
workflows in many scientific areas including
neuroinformatics and in the domain of electro-
physiological experiments.
All of the mentioned engines use the parameter
type control during data processing (Stebetak, 2013).
This simple comparison of parameters ensures that
only compatible methods can be connected.
However, methods used in the electrophysiology
domain are specific in case of syntax and semantics
for various inputs/outputs. For example, only a
subset of the result of a previous method can be used
as an input to a next method. This case is not solved
by these engines.
For well-designed workflows, ensuring syntac-
tical compatibility is necessary but not a single step.
Used methods have to be also connected correctly in
terms of their semantics. However, semantics of
piped methods (if the connection makes sense or
not) is not satisfactorily solved by these engines.
3 ANALYTIC METHODS AND
ALGORITHMS
The following subsections briefly describe a set of
methods suitable for EEG/ERP signal analysis.
These methods are used for detection of ERP
waveforms or artifact removal.
3.1 Signal Preprocessing
A pure EEG signal contains a lot of artifacts (non-
cerebral signal); ERP waveforms are hidden.
Therefore, signal preprocessing methods are used for
suppressing artifacts and obtaining ERP waveforms.
An EEG signal is divided into epochs. Each
epoch starts at the time when a stimulus appeared
and its length depends on the latency and length of
ERP waveforms. In ERP experiments, several types
of stimuli are used.
Averaging (Rondik, 2012) is a common method
for highlighting ERP waveforms. Since the
background EEG has a higher amplitude then ERP
waveforms, the averaging technique highlights the
waveforms and suppress the background EEG
(Vidal, 1977). A set of epochs is the input of the
averaging method. The output of this method is an
averaged signal belonging to a specific stimulus.
3.2 Signal Processing
We widely use the following signal processing
methods: Fast Fourier transform, Matching Pursuit,
Discrete and Continuous Wavelet transform, ICA,
and Hilbert-Huang transform (Ciniburk, et al. 2010).
This section briefly describes principles of these
algorithms.
The Fourier transform converts waveform data in
the time domain into the frequency domain. Since
artifacts usually have higher amplitude and
frequency than a normal ERP component, this
technique is useful for detecting artifacts within the
EEG or ERP signal.
The matching pursuit (MP) algorithm is
frequently used for continuous EEG processing. It
decomposes any signal into a linear expansion of
functions called atoms. An input signal is
approximated by a Gabor atom, which has the
highest scalar product with the original signal, and
then it is subtracted from the signal. This process is
repeated until the whole signal is approximated by
Gabor atoms with an acceptable error (Vareka,
2012).
Wavelet Transform (WT) (Ciniburk, et al. 2010)
is a suitable method for analyzing and processing
non-stationary signals such as EEG. For EEG signal
processing it is possible to use continuous wavelet
transform (CWT) or discrete wavelet transform
ModelofSyntacticCompatibilityinWorkflowsforElectrophysiology
443
(DWT). Both CWT and DWT were tested during
our research focused on automatic ERP detection.
DWT is common in computer science because of
high performance caused by its algorithmic
complexity. In automatic ERPs detection it is
necessary to have a wavelet which corresponds to
a detected ERP component as much as possible.
CWT is often replaced in computer science by its
discrete form because of its algorithmic complexity.
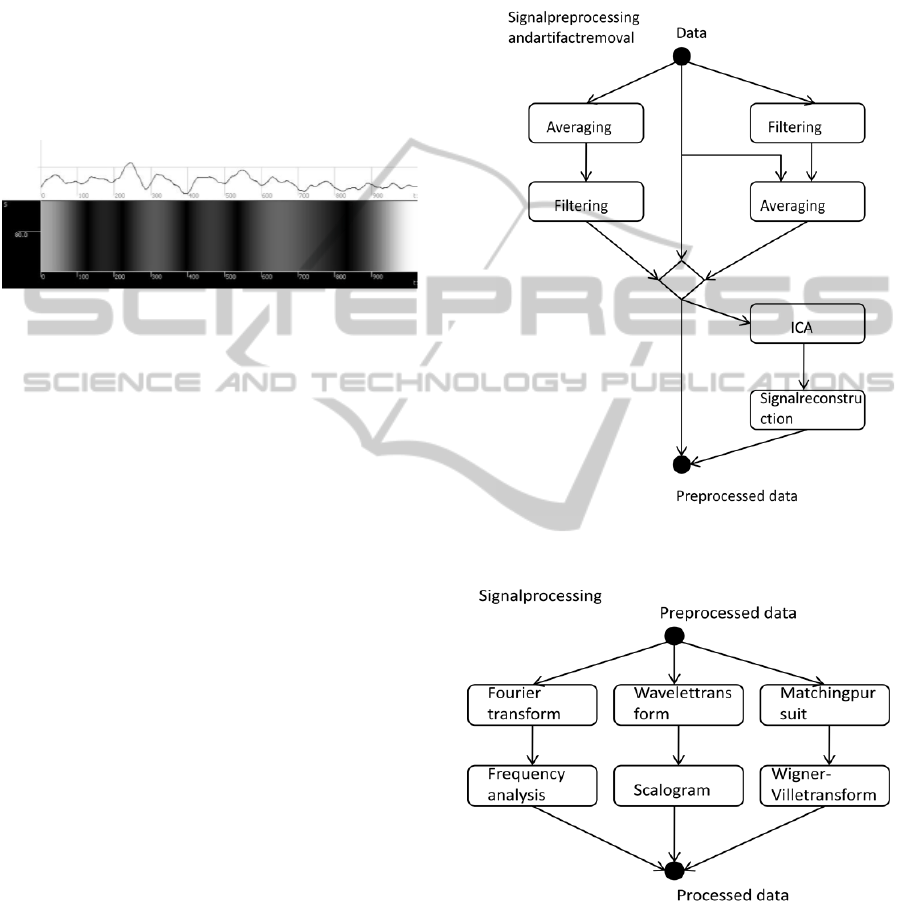
The result of the wavelet transform is visualized in a
scalogram (Figure 1).
Figure 1: Input signal and its scalogram. (Rondik, 2012).
Independent Component Analysis (ICA)
(Hyvärinen, et al. 2001) is a method for blind signal
separation and signal deconvolution. In the
EEG/ERP domain, ICA can be used for artifact
removal, ERPs detection, and – generally speaking –
for detection and separation of every signal which is
independent on EEG activity.
The Hilbert-Huang transform (HHT) was
designed to analyze nonlinear and non-stationary
signal. It can be used for detection of ERP
waveforms (Ciniburk, 2011).
4 WORKFLOWS IN
ELECTROPHYSIOLOGY
Data obtained from electrophysiological
experiments are mostly analyzed using the methods
described in Section 3. However, there is usually
a need to use more than one method for analyzing an
EEG/ERP signal. Therefore, we provide an
opportunity to define workflows for complex
analysis of experimental data.
In the mentioned domain, a workflow includes a
complex set of analytic methods that process
experimental data sequentially or in parallel.
Workflows are organized as a tree structure,
where each branch of the tree has the same meaning
as a pipe in Linux; an output of the method serves as
an input of the next method. We define steps
between methods in sequential workflows. These
steps ensure that a result from a previous method is
transferred to a next method. Since different
methods have various input/output parameter types,
we have to secure their syntactic compatibility.
In Figures 2 and 3, the preprocessing and
processing methods suitable for giving into a pipe
are shown.
Figure 2: Signal preprocessing and artifact removal
(Stebetak, 2013)
Figure 3: Signal processing (Stebetak, 2013).
Note that ensuring both syntactic and semantic
compatibility of methods is important for well-
designed workflows. This paper is focused on
presenting an innovative approach in case of
ensuring syntactic compatibility of methods in
workflows. We will focus on modelling semantic
compatibility in our future work.
HEALTHINF2014-InternationalConferenceonHealthInformatics
444
5 SYNTACTIC COMPATIBILITY
EXTENSION IN
ELECTROPHYSIOLOGY
It is necessary to ensure the syntactic compatibility
in workflows. It means that the output of a previous
method and the input to a next method must match.
Otherwise, the syntactic error will occur.
The syntactic compatibility is usually ensured by
the parameters type comparison. However, the
methods in the electrophysiology domain can return
more than one result type. It is also possible that
only a subset of result is used as an input to the next
method in a workflow. The next paragraphs describe
proposed extension of ensuring syntactic
compatibility.
5.1 Simple Comparison of Parameter
Type
Each used method has a definition of input/output
parameter types. We define these types via XML file
attached to a method. An example of input/output
parameter type of a method is given below.
<?xml version="1.0" encoding="UTF-8"?>
<method name="MP_1.0.0">
<param type="input" format="ARRAY"
datatype="DOUBLE" />
<param type="output"
format="2DARRAY" datatype="DOUBLE"
/>
</method>
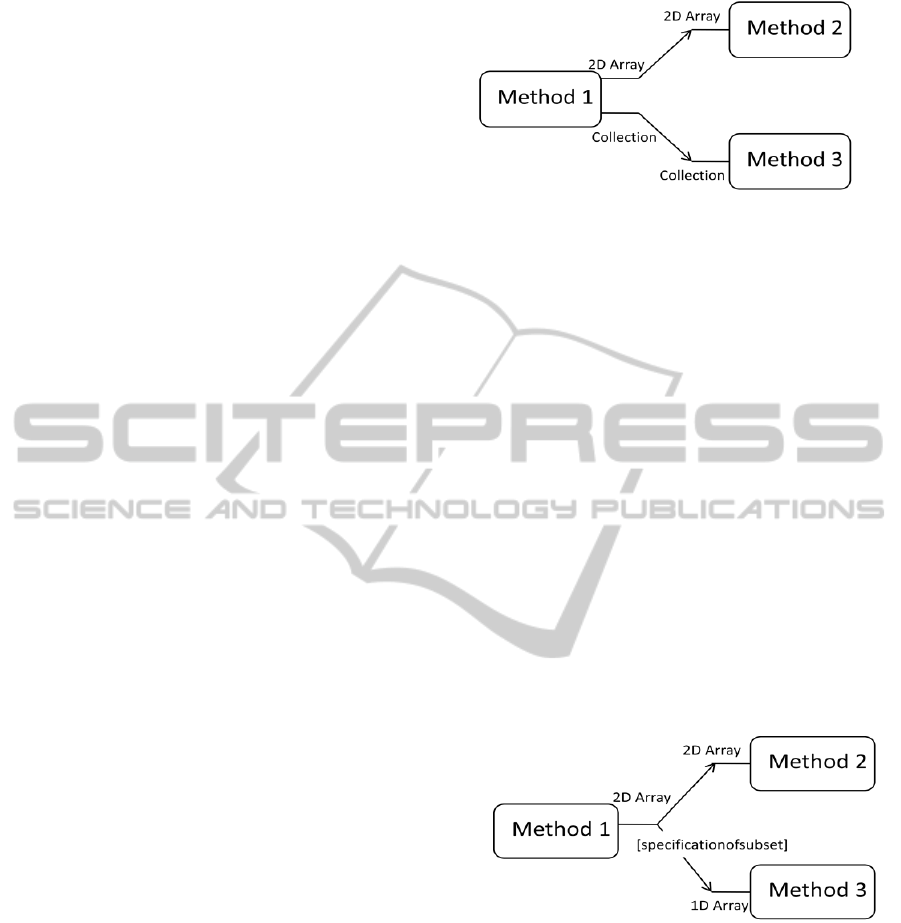
5.2 Multi-format Parameters
Because of variety of input/output formats, we
extended the implemented methods by multi-format
parameters. It means that the methods accept more
input formats and return more output formats
(Figure 4). In this example, Method 1 provides result
in format of a two-dimensional array and also in data
collections, e.g. Map in Java or Dictionary in C#.
Method 2 accepts input in two-dimensional array
format and Method 3 accepts data collections. Both
these methods can be added into a sequential
workflow following the Method 1 since this method
provides a multi-format output.
The syntactic compatibility of methods is
ensured, when one of output parameter types of
a previous method matches with an input parameter
type of a next method.
Figure 4: Multi-format output of the result provided by
Method 1.
5.3 Subset of Result
In electrophysiology, we often use methods that
provide results in a different format than a next
method requires, e.g. the method for detection of
epochs (Section 3.1). This method returns signal
belonging to all detected epochs but only signal
from one epoch for further processing (e.g.
averaging) is used.
An example of using a subset of result is given
in Figure 5. In this case, Method 1 returns results
only in a two-dimensional array format. The input of
Method 2 has a two-dimensional array as well.
Therefore, these methods are compatible by simple
comparison of their parameter types. On the
contrary, Method 3 expects a one-dimensional array
as an input. Therefore, a scientist has to select a
subset from two-dimensional array produced by
Method 1.
Figure 5: Scientist specifies a subset of result for Method
3.
When a scientist (a user in general) puts methods
such as Method 1 and Method 3 into a workflow, the
workflow processing stops and the results from
Method 1 is displayed. The user is requested to
choose a subset of the result that is used as an input
to Method 3. Then the workflow processing
continues.
ModelofSyntacticCompatibilityinWorkflowsforElectrophysiology
445

6 CONCLUSIONS
This paper summarizes methods for EEG/ERP
signal preprocessing and processing. It brings an
introduction to principles of these methods as well
as their using for ERP waveforms detection or
artifacts removal.
Since analyzing an EEG/ERP signal usually
includes using more methods sequentially or in
parallel, definition of workflows for complex
analysis is presented.
Since methods are executed sequentially, it is
necessary to ensure that the execution of workflow
does not fail due to incompatibility of piped
methods. In electrophysiology, there are methods
with various input and output formats. The proposed
solution ensures syntactic compatibility of piped
methods. It includes an extension of used methods
by multi-format parameters described in Section 5.2.
This solution also enables using a subset of a result
of a previous method as an input to a next method.
Our future work will focus on testing the
proposed solution by implementation of workflow
steps into our neuroinformatics infrastructure. We
will also focus on modelling semantic compatibility
of methods.
ACKNOWLEDGEMENTS
The work was supported by the UWB grant SGS
2013-039 Methods and Applications of Bio- and
Medical Informatics.
REFERENCES
(CARMEN) Development of a workflow system for the
CARMEN Neuroscience Portal (2013)
http://neuroinformatics2012.org/abstracts/development-of-
a-workflow-system-for-the-carmen-neuroscience-
portal
Taverna (2013), http://www.taverna.org.uk/
eScience Central (2013) http://
www.esciencecentral.co.uk/?p=151
Watson, P., Hiden, H., Woodman, S., 2010, “e-Science
Central for CARMEN: Science as a Service.”
Concurrency and Computation: Practice and
Experience, Volume 22, Issue 17, pages 2369-2380,
10 December.
Rondik, T. 2012, “Methods for Detection of ERP
Waveforms in BCI Systems” State of the Art and
Concept of Ph.D. Thesis, Pilsen.
Vidal, J. J., 1977, Real-time detection of brain events in
EEG. Proceedings of the IEEE, Volume 65, Issue 5,
pp. 633 - 641.
Ciniburk, J., Mouček, R., Mautner, P., Řondík, T., 2010,
ERP components detection using wavelet transform
and matching pursuit algorithm, DCII,Prague (2010)
Vareka, L., 2012, Matching Pursuit for P300-based Brain
Computer Interfaces, Prague.
Hyvärinen, A., Karhunen, J., and Oja, E., 2001,
“Independent Component Analysis” Adaptive and
Learning Systems for Signal Processing,
Communications and Control. J. Wiley,.
Rondik, T., 2010, “Použití matching pursuit s vlastním
slovníkem funkcí při detekci ERP v EEG signálu
(Using matching pursuit algorithm with its own
dictionary for ERP in EEG signal detection)”.
Proceedings of the 10th Conference Kognice a umělý
život (Cognition and Artificial Life), Opava: Slezská
univerzita, pp. 329-332,.
Ciniburk, J., 2011, “Hilbert-Huang transform for ERP
detection“, Ph.D. Thesis, University of West Bohemia,
Pilsen, Czech Republic.
Stebetak, J., 2013, “Analytic Methods and Workflows for
EEG/ERP Domain” State of the Art and Concept of
Ph.D. Thesis, Pilsen.
Littauer, R., Ram, K., Ludäscher, B., Michener, W.
Koskela, R., 2012, “Trends in Use of Scientific
Workflows: Insights from a PublicRepository and
Recommendations for Best Practice“, The
International Journal of Digital Curation, Volume 7,
Issue 2.
HEALTHINF2014-InternationalConferenceonHealthInformatics
446
