
A Pareto Front Approach for Feature Selection
Enguerran Grandchamp
1
, Mohamed Abadi
2
and Olivier Alata
3
1
Laboratoire LAMIA, Université des Antilles, Campus de Fouillole, 97157 Pointe-à-Pitre Guadeloupe, France
2
Institut XLIM-SIC, UMR CNRS 6172, Université de Poitiers, BP 30179, 8962 Futuroscope-Chasseneuil Cedex, France
3
Lab. Hubert Curien, UMR CNRS 5516, Univ. Jean Monnet Saint-Etienne, Univ. Lyon, 42000, Saint-Etienne, France
Keywords: Hybrid Feature Selection, Mutual Information, Multiobjective Optimization, Pareto Front, Classification.
Abstract: This article deals with the multi-objective aspect of an hybrid algorithm that we propose to solve the feature
subset selection problem. The hybrid aspect is due to the sequence of a filter and a wrapper method. The filter
method reduces the exploration space by keeping subsets having good internal properties and the wrapper
method chooses among the remaining subsets with a classification performances criterion. In the filter step,
the subsets are evaluated in a multi-objective way to ensure diversity within the subsets. The evaluation is
based on the mutual information to estimate the dependency between features and classes and the redundancy
between features within the same subset. We kept the non-dominated (Pareto optimal) subsets for the second
step. In the wrapper step, the selection is made according to the stability of the subsets regarding classification
performances during learning stage on a set of classifiers to avoid the specialization of the selected subsets
for a given classifiers. The proposed hybrid approach is experimented on a variety of reference data sets and
compared to the classical feature selection methods FSDD and mRMR. The resulting algorithm outperforms
these algorithms.
1 INTRODUCTION
Feature Selection (FS) is an active topic of interest. A
large number of algorithms have been proposed. The
basic idea is to select a subset from a large set of
features. FS is a branch of the Dimension Reduction
problem (Hilario, 2008). FS is an important task in
many fields such as text characterization, image
research, bioinformatics, color image processing,
data mining, etc. The aim is to select relevant features
for knowledge interpretation or representation,
computation time reduction and overall improvement
in performance (such as classification accuracy).
The relevancy of the features can have different
definitions depending on the application: in
knowledge interpretation or representation, the size
reduction and the semantic and/or the diversity of the
selected features are important in order to keep in a
lower dimension the topological structure of the
information; for classification applications,
relevancy is directly linked to a good rate in learning
or classification; in protein biomarkers identification,
the reduction of the feature subset size and its stability
when applying different learning sets are more
important than classification performances. The
relevancy is linked to the quality, the complexity, the
diversity or the performance of the feature subset.
Different approaches have been developed to
select a subset of features. They differ by their
research method to explore the subsets, their criterion
for comparing and ranking them and their selection
process.
We design a hybrid method to combine the
advantages of both filter and wrapper approaches: a
fast (filter) way to select diversified subsets (multi-
objective) having good internal properties (filter) and
a final selection based on performances (wrapper).
The stability criterion avoid specializing the subsets
to a given classifiers.
After a general presentation of the main
exploration methods, the fitness functions and
selection processes are presented in section 2. Section
3 presents the multi-objective principle. Then in
section 4, we present the hybrid method and the
criterions. In section 5, some formalism is given
concerning the criterion, the non-domination
principle and the algorithm. The algorithm is given in
section 6. The experiments on benchmarking
database, classification and segmentation
applications are given in section 7. Finally, section 8
gives conclusions and perspectives of the work.
334
Grandchamp, E., Abadi, M. and Alata, O.
A Pareto Front Approach for Feature Selection.
DOI: 10.5220/0005752603340342
In Proceedings of the 5th International Conference on Pattern Recognition Applications and Methods (ICPRAM 2016), pages 334-342
ISBN: 978-989-758-173-1
Copyright
c
2016 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved

2 THE FEATURE SELECTION
PROBLEM
Many papers have been published on the modeling
and the description (Somol, 2010) of feature selection
problem. We summarize the main ideas implemented
in the different feature selection approaches.
Categorization is done according to
Exploration methods (Sun, 2010) :
• Greedy methods based on sequential approaches
such as Sequential Forward Selection (SFS) and
Sequential Backward Selection (SBS).
• Sequential Forward Floating Selection (SFFS)
and Sequential Backward Floating Selection
(SBFS).
• Genetic Algorithms (GA)
Fitness functions:
• A quality measure evaluated on each features
separately: Dependency, entropy, relief-f,
distance measures, statistical measures and more
recently probabilistic measures based on the
estimation of Mutual Information (Peng, 2005);
or directly on the subset: correlation,
redundancy, Information Criteria, for example.
• A performance measure: the good classification
rate or error rate during the learning step.
• A complexity measure: the cardinality of the
subset, the complexity of the classifiers.
Selection processes:
• A single candidate selection for sequential
approaches: maximization (classification rate,
relevancy, etc.) or minimization (error rate,
correlation, etc.) of the criterion.
• Multiple candidates for evolutionary approaches
The main used evaluation is based on their
performances in a classification context.
These different approaches lead to the separation of
the methods in four families based on how to compare
and rank the subsets:
• Wrapper methods use a machine learning
algorithms during the exploration step to
evaluate the candidate’s subsets and the
corresponding classifier during the evaluation of
the returned solution (test stage). It often gives
the best performances but it is time consuming
because of the training step on classifiers.
• Filter methods use an independent criterion to
measure the quality of the feature subsets. These
methods are the most popular because they
considerably reduce the computation time.
• Embedded methods try to combine the
advantages of both approaches. Nevertheless, the
computation time still remain important.
• Hybrid methods use a sequence of Filter and
Wrapper methods (Peng, 2005).
More details are given in the previous references and
particularly in (Hilario, 2008) and (Somol, 2010)
which are surveys of methods.
3 A MULTI OBJECTIVE
APPROACH
Most of the time the exploration methods deal with a
single criterion. However the use of only one
characteristic to rank and select the subsets is
insufficient in many cases. Authors then defined
combinations of several criterions to integrate quality
and performance. In practice, defining a combination
of criterions is not an easy task. It depends on the
application and often requires parameters to balance
the different parts of the criterion. These criterions
generally have opposite behaviors because increasing
the performances often requires adding features
which increase complexity.
In order to bypass this drawback, a multi-
objective approach has been adopted in some studies
(Hasan, 2010). A multi-objective approach try to
simultaneously optimize several fitness functions
during the exploration. However the criterions often
have opposite behaviors leading to a set of non-
dominated solutions called the Pareto set.
For the FS problem, the different approaches, deal
with wrapper methods using GA as exploration
method with a simple binary encoding and standard
crossover and mutation. One of the objective is the
cardinality of the subset and the other one a
classification rate or error rate.
4 THE PROPOSED HYBRID
APPROACH
4.1 Filter and Wrapper Combination
Hybrid methods are proposed in the literature (Cantu-
Paz, 2004), (Peng, 2005), but the main objective of
these works is to reduce the computation time. Indeed
the criterion used in the wrapper step is the classifier
performance in a mono-objective approach. The filter
method is used to reduce the exploration space in a
very high dimensional data set by evaluating the
quality of the features in a mono-objective way:
Kullback-Leiber distance between histograms of
feature values; mRMR criterion; the relief criterion
and; the relative certainty gain.
A Pareto Front Approach for Feature Selection
335

The way to select the subsets for the wrapper step
represents the main differences between the
approaches. In (Cantu-Paz, 2004) they select the
features by fixing a threshold on the Kullback-Leiber
distance; In (Peng, 2005), they keep subsets having a
classification error under a given threshold.
As the number of features is reduced by the filter
step, the wrapper step manages the retained features
by the mean of a classical GA, or sequential forward
and backward search with a classification accuracy
criterion.
We propose a hybrid method by combining the
Filter and Wrapper methods in two sequential steps.
This approach improves the lack of diversity of the
solutions returned by standard algorithms and reduces
the dependency between subsets and classifiers. The
computation time remains acceptable thanks to the
use of a fast filter approach and a controlled
exploration of Pareto solutions during the first step.
These procedures coupled with a multi-objective
approach with two quality objectives allow keeping
diversity. All the selected subsets using the Pareto
front are evaluated during the wrapper step.
We prefer a stability criterion to select the final
subsets instead of raw performances regarding one
classifier, in order to keep performances and
independency between subsets and classifiers.
Indeed, we are looking for diversified subsets in
the filter step in order to have different kinds of
solutions to be evaluated during the wrapper step to
increase the probability to reach stable ones. In this
way, the building of the Pareto front seems to be the
more appropriate choice.
4.2 Criterion and Diversity
The second stage of some previous approaches
maintains a kind of diversity by the crossover step and
the mutation step of a GA. On the other hand, the
selection of the first pool of features by the filter step
is done using a single criterion which restricts the
explored subsets. Indeed, the evaluation of the subsets
is done in a single way which leads to reject subsets
having good properties according to another criterion.
This is particularly the case for single criterions
which are composed of multiple parts (mRMR for
example, composed of Redundancy and Relevance).
In this context, solution having very low redundancy
or very high relevance could be rejected by the
selection process if the resulting aggregation function
has a low evaluation. To increase the diversity of the
selected subsets our filter step explores the space in a
multi-objective way with two quality objectives and
a complexity objective.
The evaluation of the quality is based on the Mutual
Information (MI) to separately measure the
Dependency (D) and Redundancy (R) of the subsets.
The theoretical interest for Mutual Information has
been proved in (Peng, 2005). These two criterions
measure both the individual quality of the selected
features and the quality of the subset. The separate
evaluation of these two measures (contrary to (Peng,
2005)) is important because a relevant subset is not
necessarily a subset containing only significant
attributes taken alone. Indeed the relevance of a
subset may be due to combinations of features.
5 CRITERIONS
The criterions are based on the mutual information
which is considered to be a good indicator to study
the dependency between a feature and the
classification and the redundancy between random
features.
Mutual Information
Let and be two random variables with discrete
probability laws. The Mutual Information (MI)
(
;
)
is defined by
(
)
,
(
)
and
(
,
)
.
(
;
)
=
(
,
)
∙log
(
,
)
(
)
∙
(
)
∈
∈
(1)
with Ω
X
and Ω
Y
the sample spaces of X and Y
respectively.
When and are dependent,
(
;
)
is high.
(
;
)
is equal to zero when and are
independent.
Selection criterion definition
For each subset of features, we define the
relevance expressed by the Dependency (D) which is
the average MI between the variables of S (X
i
) taken
separately and the class of the samples modeled by a
discrete random variable called c with sample space
equal to the class labels:
=
|
|
∑
(
;
)
∈
(2)
(
;
)
represents the MI between a variable and the
classes. It translates how X
i
is useful to describe the
classes.
The Dependency has to be maximized. However
in order to have a homogenous expression of the
objective we prefer to express the opposite of the
Dependency (-D) to minimize each criterion.
The feature selection using only is not optimal
because of redundancy between the variables.
ICPRAM 2016 - International Conference on Pattern Recognition Applications and Methods
336
Different ways exist to measure the redundancy and
we use the one expressed in (Peng, 2005). It is based
on the computation of the average MI between two
variables
;
,,⋯,
belonging to the same
subset having variables.
=
|
|
∑
;
,
∈
(3)
The redundancy must be minimized.
6 HYBRID AND
MULTIOBJECTIVE
ALGORITHM
6.1 Principle
The novelty of our approach is that previous
criterions are treated separately contrary to (Peng,
2005) and (Al-Ani, 2002) where criterions are
combined to produce the mRMR (minimal-
redondance-maximale-pertinence) criterions (ex.
max
(
−
)
or max
(
/
)
). These mono-objective
criterions didn’t ensure the simultaneous
convergence of criterions
(
2
)
and
(
3
)
to their optimal
value but lead to a trade-off between them.
We also keep the subset cardinality (L) which must
be minimized as a third criterion.
The goal of a multi-objective optimization is to
improve several criterions. When these criterions
have opposite behaviors considering the research of a
solution, we necessarily have to degrade at least one
criterion to improve another one. This leads to
different kind of solutions which are not necessarily
comparable. If we don’t want to make a choice
between solutions we must keep all solutions being
better than any others on at least one criterion. This
leads to the notion of domination which is essential to
ensure diversity in the final sets.
Without loss of generality, we illustrate this notion
in our particular case.
Following the previous section, each subset is
evaluated with three values (
,
,
)=(−D,R,L).
• A subset S dominates a subset S
2
according to
if
(
S
)
<
(
S
)
. i=1, 2, or 3
• A subset S dominates a subset S
2
if ∀
(
S
)
≤
(
S
)
and ∃|
(
S
)
<
(
)
.
• A subset S is not dominated if
∄
|
dominatesS (∄
|∀
(
)
≤
(
S
)
, ∃
(
)
<
(
S
)
).
• The set of all non dominated subsets is called the
Pareto set.
The third criterion, which represents the
complexity of the subset through its cardinality,
allows keeping subsets with different size (for low
number of features the redundancy may be better and
for high number of features the dependency may be
better). Nevertheless, even if we have one Pareto
front for each possible subset size, there is no
certainty to obtain at least one subset for each possible
size. This could be an inconvenient for some
applications. In such condition, the exploration step
can deal with only the quality criterions and each
intermediate Pareto front (corresponding to a specific
size) could be kept. This approach called Multi Pareto
Front (MF) is detailed in the next paragraph.
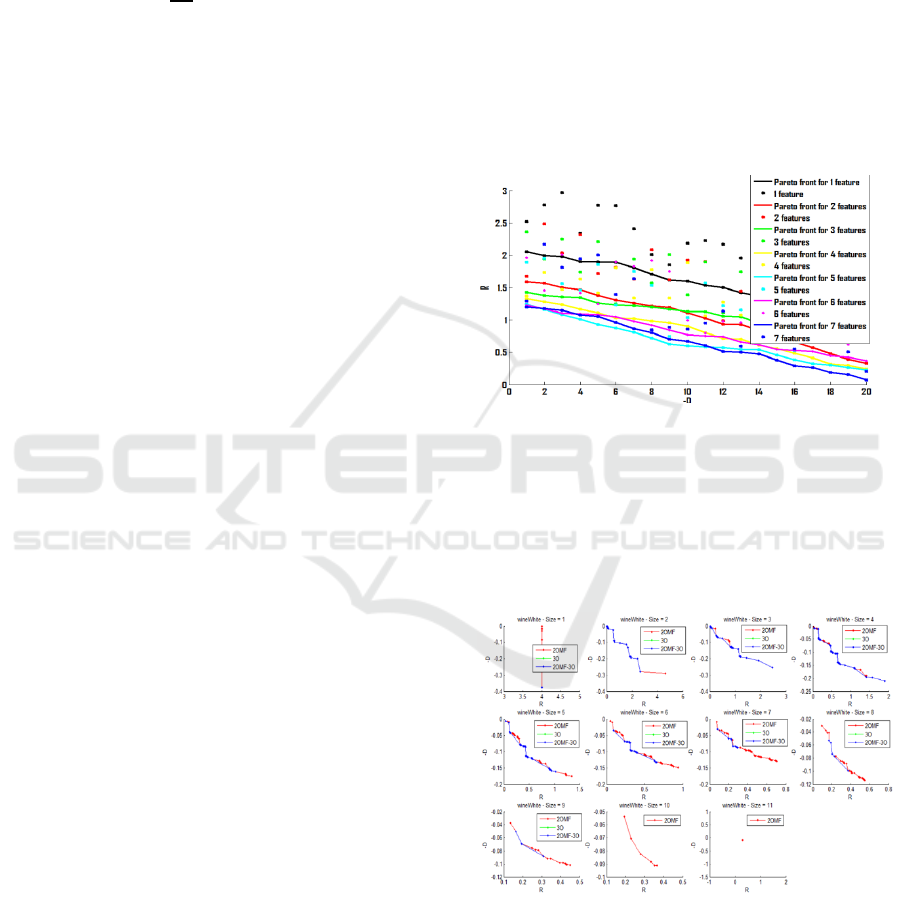
Figure 1: Pareto front evolution principle.
Figure 1 illustrate the evolution of the Pareto front
projected in (-D,R) space. As the number of features
increases the solutions in the Pareto front tend to
decrease –D and R values. Figure 2 shows the same
with in a real case using wineWhite database of the
UCI.
Figure 2: Pareto front evolution for wineWhite UCI
database.
6.2 Filter Step: Multiobjective
Exploration
For any optimization problem, a unique Pareto set
exists for a given data set and the considered
A Pareto Front Approach for Feature Selection
337

criterions. In a muti-objective context, an exhaustive
search, or an algorithm having asymptotic
convergence properties such as Genetic algorithm, is
classically required to find this set. Both are time
consuming and sometimes too slow to reach the
optimal Pareto set in a reasonable time. In practice,
people build a sub-optimal Pareto front which is the
Pareto front computed over the visited solutions. One
of the main qualities of a search method is then its
ability to provide solutions close to the ones of the
optimal Pareto front. Our filter search method joins
this way and has been developed to approach the
building of the optimal Pareto front.
The filter step uses a sequential forward search to
explore the subset space adopting the following
algorithm:
1. Let =
|
∈ [1,]} be the complete set
2. We start with all possible pairs of
features
=
,
∈
[
1,
]
,∈
[
1,
]
,≠}.
3. Each subset S is evaluated with (-D(S),
R(S)) criterions and the non-dominated
subsets (
) are preserved.
is the
Pareto front at iteration 2 (
=
).
4. At iteration k,
is the non dominated
subsets of size k (k>2) and
is the
global Pareto Front (
=
⋃
).
5. We build V
k+1
by adding to
one new
feature taken within the remaining
features:
=
(
∪
)|
∈
,
∈\}.
Each subset S in V
k+1
is then evaluated
with (-D(S), R(S)).
6. We build ND
k+1
by retaining the non
dominated subsets of size k+1 within
.
We note that ND
k+1
⊆
. This step is
required because some subsets of
can
be dominated by ones of
(opposite is
impossible because each subset in
is
greater than the ones in
).
7. The algorithm ends if k=M.
This algorithm is called two Objectives Multi-
Front Algorithm (2OMF) and the returned set of
subsets is
=
⋃
.
We can note that
⊆
. Indeed, subsets of
couldn’t dominate subsets of
because these
last ones have a lower size: ∀
∈
,
(
)
≤
,∀
∈
,
(
)
=+1.
6.3 Wrapper Step: Stability Criterion
The wrapper step is used to rank the selected subsets
and to select a subset considering the application. For
this step, the exploration space has been sufficiently
reduced during the filter step to allow an exhaustive
evaluation of the remaining subsets ND
F
. A large
majority of wrapper approaches deals with Feature
Selection in terms of performances regarding a
classifier, but few studies select subsets for their
stability. Nevertheless, the stability is a topic of
interest in studies dealing with high dimensional data
and a small number of samples (Hilario, 2008).
Moreover, wrapper methods can lead to good
classification accuracy for a specific classifier but
with poor generalization properties (Kalousis, 2007),
(Peng, 2005) (i.e. over-fitting for one classifier and
low performances for another one).
The stability is defined by (Somol, 2010) as being
the quality of a subset to have the same performances
with different training sets. Different stability indices
can be used such as Hamming distance, correlation
coefficients, Tanimoto distance, consistency index
(simple, weighted or relative weighted) and Shannon
entropy. In (Kuncheva, 2007) and, the stability is
measured by running a wrapper scheme several times
with a unique classifier and different learning sets (no
cross-validation). The stability is based on an
evaluation of the similarity between subsets returned
by different runs. If the index is high the subset is
selected. Otherwise the selection is based on the
classification rate evaluation.
In this paper, we investigate another kind of
stability between different classifiers (each trained
and evaluated with a cross-validation process).
According to this kind of stability has been neglected
in the literature. A subset is stable regarding
classifiers if the performances obtained with different
classifiers are close. The easiest way to compute the
stability of a subset is to compute the amplitude of the
classification rates obtained with several classifiers
K-Nearest Neighbor (KNN), Linear Discriminant
Analysis (LDA), Mahalanobis (Mah), Naive Bayes
(NB), Simple Vector Machine (SVM) and
Probabilistic Neural Network (PNN):
() =max
∈
R
(S)
}
−min
∈
R
(S)
}
(4)
Where S is the subset, Cl a set of classifiers and R
(S)
the classification rate obtained with the classifier c
applied on the subset S.
Finally, we identify the stable and successful
subsets. Therefore, the selection of the interesting
subsets is done in a two objectives way by
maximizing the mean classification rate (()) and
by minimizing the amplitude (()).
()=mean
∈
R
(S)
}
(5)
7 RESULTS
In a first step we present the results obtained with the
2OMF algorithm. In a second step we compare the
ICPRAM 2016 - International Conference on Pattern Recognition Applications and Methods
338
2OMF method with two other existing feature
selection methods: mRMR (Peng, 2005) and FSDD
(Liang, 2008). Both are using filter criterions to select
the features and then they evaluate the unique
returned solution using classifiers. We choose mRMR
method because it uses the same criterions as 2OMF
but in a mono-objective way. We choose FSDD
because it is a fast algorithm which converges to the
optimal solution regarding a distance criterion. In
both cases, it is interesting to project the solutions
obtained with different filter steps in the space
(performance, stability) of the wrapper step and to
compare them with our pool of solutions. The
comparison is done by means of the size and the
stability of the subsets returned by each method and
also the computational time of each method.
Each step uses UCI databases for validation and
more particularly iris, TAE, abalone,
PimaIndiansDiabetes, wineRed, wineWhite, wine,
imgSeg, ionosphere and landSat databases containing
4, 5, 7, 8, 11, 11, 13, 18, 34 and 36 features
respectively. Figure 3 to 5 present some of the
obtained results. The stability is computed after
applying KNN, LDA, Mah, NB, and PNN classifiers.
7.1 2OMF Method Analysis
We analyse more precisely the results of the 2OMF
algorithm after the second step of the algorithm. This
step is based on the wrapper approach which sorts
and then selects among the retained subsets during the
filter step. We recall that the used criterion is the
stability (in a two objectives way) when different
classifiers are applied. In this space (mean rate,
amplitude) we compute a new Pareto front composed
of several solutions and we focus on them to select
the most interesting ones.
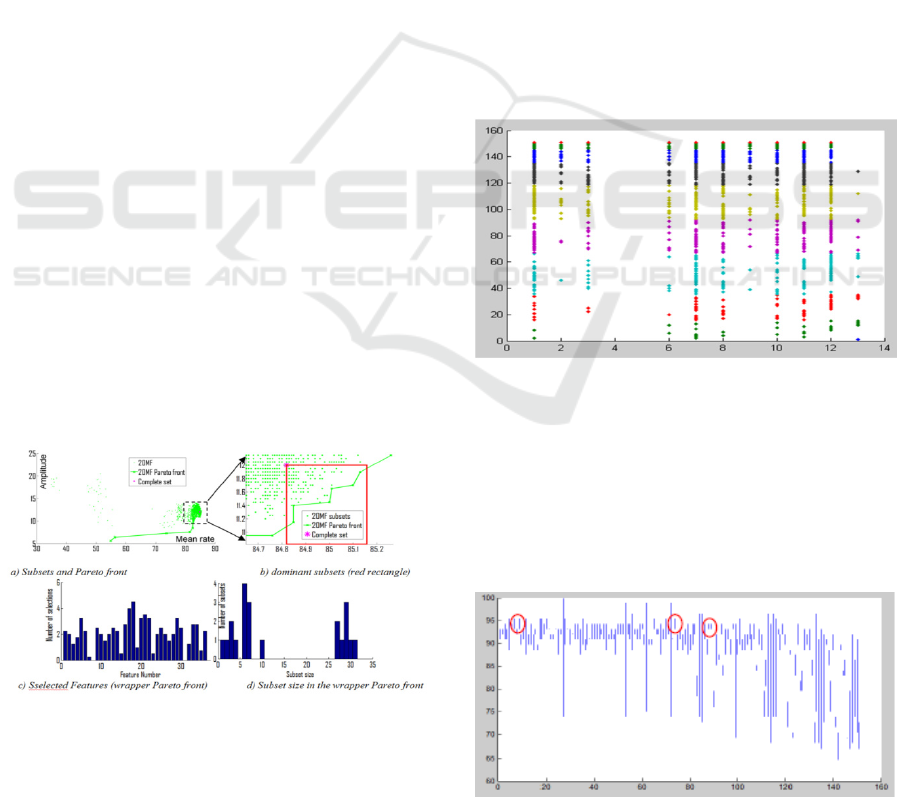
Figure 3: Stability analysis for landSat database.
The Figure 3 displays information about the
stability of the selected subsets after filter step (green
points) for landSat database. As showed in Figure 3
b) (which is a zoomed part of the Figure 3 a) ), a lot
of subsets dominates the complete set (purple star in
the figure) even if they are not in the Pareto front:
these subsets are within the red rectangle. All of these
subsets have higher mean classification rate and
lower amplitude than the complete set. They can also
be interesting because some of them have lower
number of features than the one in the Pareto Front
and a quite good classification stability as it is better
than the complete set stability. For the studied
database, there are 21 subsets in the front (6
dominating complete set) and 73 subsets that
dominate the complete set.
Figure 3 c) shows the histogram of the selected
features computed using all subsets of the 2OMF
wrapper Pareto Front. We note that quite every
features are represented. In the same way Figure 3 d)
gives the repartition of the size of the Pareto subsets
in order to illustrate the diversity of the solutions.
Landsat database is composed of 36 features and
some of the subsets in the Pareto front are composed
of less than 10 features. A further analysis of the
subset sizes is given in the next section.
Figure 4: Wine features used.
Figure 4 shows the features used by the different
Pareto solutions for the wine database. We remark
that some features are not present (4 and 5) or
underrepresented (2, 9) in the Pareto front and some
are overrepresented (1, 7, 8, 11, 12). This suggests a
individual quality evaluation of the features which
could be studied later.
Figure 5: Wine Pareto front wrapper stability.
A Pareto Front Approach for Feature Selection
339

Figure 5. shows the min and max good classification
rate after the wrapper step for each Pareto front
solution for wine database. We remark a great
diversity both in terms of good classification rate than
in terms of standard deviation. In this view we are
looking for solutions having a high classification rate
and a low standard deviation (examples are given
within the red circles).
7.2 Comparison between 2OMF,
mRMR and FSDD
We compare our algorithm with two well-known
feature selection methods: mRMR and FSDD. Figure
6 displays the visited subsets using mRMR algorithm
(blue) with their corresponding Pareto front (blue
line) and using FSDD algorithm (black) with their
corresponding Pareto front (black line). We observe
that these subsets are not Pareto optimal when
compared to the 2OMF subsets (green points).
Moreover, few of them dominate the complete subset.
The same observation can be done for most of the
databases. Indeed, in few databases we observe an
mRMR subset that dominates the complete subset. A
subset of mRMR and a subset of FSDD fall into the
Pareto set only for TAE database and for iris.
Figure 6: Comparison of the stability Pareto Fronts for
2OMF (green), mRMR (blue) and FSDD (black)
algorithms.
The size of the corresponding subsets is also
displayed near the subset as well as the complete set
(purple star). We can observe that the subset size
follows high variations: between 2 and 33 for the
2OMF Pareto front for ionosphere database and
between 4 and 29 for the 2OMF Pareto front of the
landSat database for example. The mean
classification rate is also varying in a wide range: for
Table 1: Good classification rate and stability of interesting subsets (Bold faces indicate the best result(s)).
Classifiers Rate
Db Method Size Mean Var KNN Mah NB PNN
Wine
Fill set 13 87.7 29.5 70.4 92.0 96.6 79.5
Best mRMR 6 88.63 25.0 73.8 97.7 97.7 75
Best FSDD 6 88.18 25.0 73.8 97.7 95.4 75.0
2OMF
3 96.2 4.5 97.7 96.6 97.7 95.4
7 94.5 2.2 95.5 93.2 95.5 95.5
2 94.3 5.6 96.6 94.3 95.5 94.3
imgSeg
Full set 18 92.3 8.2 95.5 NA 87.3 95.3
Best mRMR 16 91.57 7.9 94.9 NA 87 94
Best FSDD 14 92.46 7.87 95.3 NA 87.5 95.4
2OMF
7 93.2 6.5 96.1 NA 89.5 96.0
8 93.1 5.9 96.0 NA 90.0 95.6
9 92.7 5.5 95.2 NA 89.8 95.4
landSat
Full set 36 84.8 12 89.4 81.6 78.5 90.5
Best mRMR 27 84.58 11.5 89.8 80.1 79.2 90.7
Best FSDD 28 84.73 11.3 89.7 80.8 79.2 90.5
2OMF
29 85.0 11.4 89.8 82.1 78.9 90.3
31 85.1 11.9 89.6 83.0 78.7 90.6
27 84.85 11.1 89.9 81.4 79.2 90.3
25 84.9 11.9 89.3 82.7 78.7 90.6
26 84.8 11.7 89.6 82.3 78.6 90.6
7 83.13 10.7 84.9 83.1 72.2 87.9
ICPRAM 2016 - International Conference on Pattern Recognition Applications and Methods
340

ionosphere database the mean rate is about 75.5% for
a two features subset, 86% for a 5 features subset and
84% for a 10 features subset; for the landSat database
the mean rate is about 82.5% for a 4 features subset
and 85.5% for a 28 features subset.
We now focus on the good classification rate
obtained for some interesting subsets. The table 1.
shows subsets obtained with 2OMF, mRMR and
FSDD methods. Information about the Pareto
optimality is also given (Underlined size in table). In
addition to this, we present whether a subset
dominates the complete set (Green background color
in table). The mRMR and FSDD subsets are chosen
among the visited ones according to their mean rate
value.
All the displayed subsets obtained with the 2OMF
method are interesting because they have a low
number of features and a better stability than the
complete set. Nevertheless, some subsets have lowest
number of features and others highest classification
rates. For example, for the wine database, a subset
with two features (features 2 and 7) have a higher
mean rate and a lower amplitude than the complete
set having 13 features. Moreover, it has a higher
classification rates for 4 classifiers over 5. In the same
way, for imgSeg database the number of features is
divided by 2 with the 2OMF method.
Let us consider now the methods from the
literature. For landSat database none of the visited
subsets dominate the complete set for both mRMR
and FSDD. Moreover, stable and successful subsets
obtained with FSDD have a higher number of features
than the ones obtained with 2OMF. Only one stable
subset having low number of features is obtained with
mRMR (8 features). However, it is dominated by the
subset returned by 2OMF which has seven features
(last line in the table). We always found a subset
among 2OMF subsets having a lower number of
features, a higher classification mean rate and a lower
classification amplitude than the best subsets returned
by mRMR and FSD.
8 CONCLUSION
This paper presents a two steps algorithm for feature
selection and studies its multi-objective aspect. The
algorithm begins with a filter step to quickly select a
first pool of subsets in a Multi-Objectives and Multi-
Fronts way (2OMF). The subsets are evaluated using
the Dependency (D) and the Redundancy (R) of the
features. Then a second step based on a wrapper
approach is applied to measure the performances of
the subsets regarding several classifiers (KNN, LDA,
Mah, NB, PNN). Then the selection of the interesting
subsets is performed using the stability of the subsets
which is evaluated with the mean and amplitude of
the classification rates. From our experimentations, it
is observed that the interesting subsets dominate the
complete set regarding both objectives. The use of the
stability to select the subsets leads to robust results
which are very interesting for some applications such
as in biology where the stability of the subsets is more
important than its raw classification rate. The
wrapper step is required because some subsets of the
filter Pareto front could have a higher classification
rate than the complete set for a given classifiers but
not for another one. A selection of features only based
on a filter method does not ensure that the selected
subset will improve classification rates for a large set
of classifiers.
The results are very convincing for all tested
databases. The subsets obtained after applying our
algorithm have lower number of features and better
classification performances compare to the complete
set of features. Moreover, the diversity of the final
pool of subsets allows selecting a subset adapted to a
specific application (good classification expected or
reduction of a high number of features). We also
compared the proposed algorithm with two feature-
selection methods (mRMR and FSDD). It is observed
that our method outperforms the other tested methods
in almost all cases.
REFERENCES
A. Al-Ani, M. Deriche, and J. Chebil, "A new mutual
information based measure for feature selection,"
Intelligent Data Analysis, vol. 7, no. 1, pp. 43-57, 2003.
E. Cantu-Paz, "Feature Subset Selection,Class
Separability, and Genetic Algorithms," in Genetic and
Evolutionary Computation, 2004, pp. 959-970.
K. Deb, Multi-Objective Optimization Using Evolutionary
Algorithms.: John Wiley and Sons, Chichester, 2001.
C. Emmanouilidis, A. Hunter, and J. MacIntyre, "A
Multiobjective Evolutionary Setting for Feature
Selection and a Commonality-Based Crossover
Operator," in Congress on Evolutionary Computation,
California, July 2000, pp. 309-316.
B.A.S. Hasan, J.Q. Gan, and Z. Qingfu, "Multi-objective
evolutionary methods for channel selection in Brain-
Computer Interfaces: Some preliminary experimental
results," in IEEE Congress Evolutionary Computation,
Barcelona, Spain, July 2010, pp. 1-6.
M. Hilario and A. Kalousis, "Approaches to dimensionality
reduction in proteomic biomarker studies," Briefings in
Bioinformatics, vol. 9, no. 2, pp. 102-118, 2008.
A. Kalousis, J. Prados, and M. Hilario, "Stability of feature
selection algorithms: A study on high-dimensional
A Pareto Front Approach for Feature Selection
341

spaces," Knowledge and Information Systems, vol. 12,
no. 1, pp. 95-116, 2007.
I. Kononenko, "Estimating attributes: Analysis and
extensions of RELIEF," in ECML-94, 1994, pp. 171-
182.
L.I. Kuncheva, "A stability index for feature selection," in
Proceedings of the 25th conference on Proceedings of
the 25th IASTED International Multi-Conference:
artificial intelligence and applications, Innsbruck,
Austria, February 2007, pp. 390-395.
N. Kwak and C.H. Choi, "Input Feature Selection by
Mutual Information Based on Parzen Window," IEEE
Trans. Pattern Anal. Mach. Intell, vol. 24, no. 12, pp.
1667-1671, 2002.
J. Liang, S. Yang, and A-C. Winstanley, "Invariant optimal
feature selection: A distance discriminant and feature
ranking based solution," Pattern Recognition, vol. 41,
no. 5, pp. 1429-1439, 2008.
E. Parzen, "On Estimation of a Probability Density
Function and Mode," The Annals of Mathematical
Statistics, vol. 33, no. 3, pp. 1065-1076, 1962.
H. Peng, F. Long, and C. Ding, "Feature selection based on
mutual information criteria of max-dependency, max-
relevance, and min-redundancy," IEEE Transactions
on Pattern Analysis and Machine Intelligence, vol. 27,
no. 8, pp. 1226-1238, 2005.
P. Somol, J. Novovicova, and P. Pudil, "Efficient Feature
Subset Selection and Subset Size Optimization," in
InTech-Open Access Publisher, vol. 56, 2010, pp. 1-24.
Y. Sun, S. Todorovic, and S. Goodison, "Local-Learning-
Based Feature Selection for High-Dimensional Data
Analysis," IEEE Transactions on Pattern Analysis and
Machine Intelligence, vol. 32, no. 9, pp. 1610-1626,
2010.
L. Zhuo, J.and al., "A Genetic Algorithm based Wrapper
Feature selection method for Classification of
Hyperspectral Images using Support Vector Machine,"
in The International Archives of the Photogrammetry,
Remote Sensing and Spatial Information Sciences, Part
B7, vol. XXXVII, Beijing, China, 2008, pp. 397-402.
ICPRAM 2016 - International Conference on Pattern Recognition Applications and Methods
342
