
C-LACE: Computational Model to Predict 30-Day
Post-Hospitalization Mortality
Janusz Wojtusiak, Eman Elashkar and Reyhaneh Mogharab Nia
Health Informatics Program, George Mason University, 4400 University Dr., MSN 1J3, 22030, Fairfax, VA, U.S.A.
Keywords: Mortality Prediction, Machine Learning, Online Calculator.
Abstract: This paper describes a machine learning approach to creation of computational model for predicting 30-day
post hospital discharge mortality. The Computational Length of stay, Acuity, Comorbidities and Emergency
visits (C-LACE) is an attempt to improve accuracy of popular LACE model frequently used in hospital setting.
The model has been constructed and tested using MIMIC III data. The model accuracy (AUC) on testing data
is 0.74. A simplified, user-oriented version of the model (Minimum C-LACE) based on 20-most important
mortality indicators achieves practically identical accuracy to full C-LACE based on 308 variables. The focus
of this paper is on detailed analysis of the models and their performance. The model is also available in the
form of online calculator.
1 INTRODUCTION
Risk Adjusted Mortality Rates are important
indicators for care outcome. They are used by
administrators, Policy makers and organizations
including government agencies, managed care
companies and consumer groups (Inouye et al, 1998)
to compare effectiveness of care among different
facilities and utilize results in quality improvement
efforts. Clinicians are mostly interested in accurate
and valid mortality prediction models to use as tools
for better planning of care, evaluation of medical
effectiveness among treatment groups while
controlling for patients’ baseline risk, and to help
clinicians decide if a patient may benefit from
intensive care units and when. From patient’s family
perspective, discussing outcome of critically ill
patients is always welcomed and appreciated.
(Rocker et al, 2004)
Many illness severity scoring systems that are
primarily used to measure prognosis early in the
course of critical illness had been widely used to
calculate in-hospital mortality. The Simplified Acute
Physiology Score (SAPS) and the Mortality
Prediction Model (MPM) use data collected within
one hour of ICU admission. Sequential Organ Failure
Assessment (SOFA) scoring uses data obtained 24
hours after admission and then every 48 hours.
Logistic Organ Dysfunction Score (LODS) and
Multiple Organ Dysfunction Score (MODS) also had
been used to measures severity of illness at time of
ICU admission. Acute Physiologic and Chronic
Health Evaluation (APACHE) scoring system widely
used to predict risk of in-hospital mortality among
ICU patients. The score uses the worst physiologic
values measured within 24 hours of admission to the
ICU and requires a large number of clinical variables
including age, diagnosis, some laboratory results, and
other clinical variables and run the result on a
computer generated logistic regression model to
calculate risk of mortality. However, these scoring
systems have shown limited accuracy predicting risk
of mortality for individual patients.
Most relevant to the presented work, the LACE
index, which has been used to predict mortality within
30 days of hospital discharge can use both primary
and administrative data. The name LACE explains
variables required: length of stay (“L”); acuity of the
admission (“A”); comorbidity or diagnoses of the
patient (uses Charlson comorbidity score) (“C”); and
number of emergency department visits in the six
months before admission (“E”). LACE index scoring
ranges from 0 (2.0% expected risk) to 19 (43.7%
expected risk) (Walraven et al, 2010). However,
standard LACE didn’t show sufficient accuracy and
it is not always possible to obtain data on the 4th item
(”E”), as emergency room visits are not necessarily
available.
Wojtusiak J., Elashkar E. and Mogharab Nia R.
C-Lace: Computational Model to Predict 30-Day Post-Hospitalization Mortality.
DOI: 10.5220/0006173901690177
In Proceedings of the 10th International Joint Conference on Biomedical Engineering Systems and Technologies (BIOSTEC 2017), pages 169-177
ISBN: 978-989-758-213-4
Copyright
c
2017 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved
169

A recent study added an extension of the LACE
(LACE+) which uses the same 4 items of LACE as
well as age and items unique to Canadian
administrative databases (such as the Canadian
Institute for Health Information Case Mix Groupings
and number of hospital days awaiting alternate level
of care arrangements). LACE+ had shown more
accuracy in predicting death within 30 days of
hospital discharge (c-statistic 0.77) than LACE index
had shown (c-statistic 0.68) (Walraven et al, 2010).
However, both instruments didn’t show sufficient
accuracy, besides it is not always possible to obtain
data on the 4th item of LACE (”E”), as emergency
room visits are not necessarily recorded in available
data.
In the presented work we propose a computational
alternative to LACE index, called C-LACE,
constructed by application of machine learning
methods to data containing information about length
of stay, acuity of the admission, and comorbidities
present during hospitalization. We decided not to use
patients’ emergency visits due to possible problems
with data availability when applying model.
A number of other models based on machine
learning and computational methods have been
proposed to predict patient mortality. For example,
(Levy et al., 2015) proposed a Multimorbidity Index
tuned to predict mortality among nursing home
patients. A number of methods have been created for
prediction of mortality among specific disease groups
such as pneumonia (Cooper et al., 1997), prostate
cancer (Ngufor et al, 2014), or sepsis (Taylor et al.,
2016).
The main contributions of the presented work are
construction of C-LACE model that can be used to
predict 30-day post-hospitalization mortality, and
more importantly detailed analysis of the model and
its behavior on real and simulated data.
2 DATA ANALYSIS AND MODEL
CONSTRUCTION
2.1 MIMIC III Data
In order to construct and test the C-LACE model, we
obtained and analyzed Medical Information Mart for
Intensive Care III (MIMIC III) data. The data is
publically available to researchers who satisfy certain
conditions (Goldberg et al, 2000). The MIMIC III
data has been collected between 2001 and 2012 in the
Beth Israel Deaconess Medical Center. It consists of
over 58,000 hospital admissions for more than 40,000
patients. It is structured into 26 tables organized as a
relational database (Johnson et al, 2016).
From the MIMIC III data, we selected only
admissions for patients at least 65 years old and alive
at hospital discharge. This results in selection of
21,651 admissions. The distribution of selected
attributes in the data is presented in Tables 1a and 1b.
The tables also show likelihood ratios (RL) associated
with each of the attributes for predicting mortality.
Within the data, the majority of patients were treated
in Medical Intensive Care Units (MICU), followed by
Cardiac Surgery Recovery Units (SCRU), Cardiac
Care Units (CCU), Surgical Intensive Care Units
(SICU) and Trauma Surgical Intensive Care Units
(TSICU). It can also be noted from the data that the
majority of patients were hospitalized only once.
In the presented work, instead of loading to
relational database, the data has been analyzed within
distributed computing infrastructure designed and
implemented as a part of the larger research project
conducted in GMU’s Machine Learning and
Inference Laboratory. The data has been mapped to
concepts within the Unified Medical Language
System (UMLS) and integrated during analysis based
on unique concept identifiers. The mapping process
is a combination of manual labor-intensive
identification of appropriate concepts which requires
strong domain background of the person performing
the mapping, with automated search for concepts
between different terminologies in UMLS. The latter
can be done when original data stored in database are
coded using one of standard terminologies, but the
final results still need to be verified by human experts.
In fact, the presented construction of the model served
as a testing application for the developed platform,
whose description is out of scope of this paper
(Wojtusiak et al., 2016).
Table 1a: Distribution of values in the data.
Died in 30 days Not died in 30 days
N = 1425 N = 20226
Age (mean, SD) 79.33 years (7.26) 76.93 years (7.16)
Length of Stay
Hospital
13.73 days (11.33) 10.52 days (9.15)
CCU (mean, SD)
121.22 days (115.56) 19.79% 72.45 days (86.18) 19.02% 1.05
CSRU (mean, SD)
262.05 days (322.26) 10.74% 92.67 days (132.29) 27.16% 0.32
MICU (mean, SD)
106.10 days (122.87) 57.89% 85.32 days (119.07) 36.14% 2.43
SICU (mean, SD)
143.88 days (222.66) 17.54% 111.51 days (170.28) 16.64% 1.07
Admission Location
Emergency Room Admit
53.75% 39.22% 1.80
Clinic Referral/Premature
18.95% 19.93% 0.94
Phys Referral/Normal Deli
6.95% 21.73% 0.27
Transfer From Hosp/Extram
18.04% 18.39% 0.98
Transfer From Skilled Nur
1.75% 0.61% 2.89
Transfer From Other Healt
0.49% 0.10% 4.75
Info Not Available
0.07% 0.00% 14.20
Variable
LR
HEALTHINF 2017 - 10th International Conference on Health Informatics
170

Table 1b: Distribution of values in the data, cont.
2.2 Model Construction
During the analysis, the data has been randomly split
into training set (80%) and testing set (20%). The
testing portion of the data has been set aside and the
experimental work has been performed on the
training set. Only final application of models has been
done on the testing set.
The data (diagnoses, ICU stays, lab tests, and
medications) has been aggregated on the level of
admission, i.e., one example in the final dataset
corresponds to hospital admission. Because of
specific implementation of machine learning library
that was used, all data had to be coded as numeric
attributes. Values of nominal attributes were coded as
0, 1, 2, etc.
- Basic demographic information (age, gender, race,
etc.) for patient has been retrieved and coded.
- Diagnoses present during hospitalization were
coded in the original data as ICD-9-CM codes. They
were aggregated to CCS categories that group
together similar ICD codes while preserving their
clinical meaning (AHRQ, 2016).
- Lab values were coded as normal and abnormal.
This coding was created as part of the original
MIMIC dataset. Then, if at least one abnormal value
for a test was detected, the overall value was coded as
abnormal. This corresponds to taking the worst case
and is consistent with several other approached to
patient modeling. However, this is a significant
oversimplification, since the values should be treated
as a time series and patient trajectory analyzed
accordingly Verduijn et al., 2007; Moskovitch and
Shahar, 2015).
- Drugs were coded with a single binary attribute
indicating use of immunosuppressant drugs. The
drugs were extracted using their LOINC codes.
- Binary output attribute indicating mortality within
30 days after discharge has been calculated using the
dates of discharge and death.
The data has been transformed into a single
analytic file (or technically corresponding data
structures) in order to be used by machine learning
software.
A number of supervised machine learning
methods have been explored in order to arrive at most
accurate and useful set of models. Among the tested
methods were logistic regression, random forest,
naïve Bayes, and support vector machines.
Comparison of the methods is presented in section
3.1, and actual descriptions of the methods is outside
of the scope of this paper and can be found in the
literature.
2.3 Implementation
The presented work has been implemented in Python
3 programming language (Anaconda distribution
Python 3.5.2). The main libraries used are Pandas (v.
0.18.1) for data processing and sciencekit-learn
(sklearn v. 0.17.1) for machine learning.
All developed source code is open source and
available on request. We are in the process of
preparing release code that will be available on the
project website.
3 RESULTS
3.1 Method Selection
The first set of results concern selection of the most
appropriate method that can handle the data. Table 2
shows comparison of accuracy of six methods applied
to training data and testing data. The methods have
been executed with multiple parameters and top
results are presented.
Table 2: Comparison of Methods applied to complete
dataset.
Method
AUC
(training)
AUC (testing)
Logistic
0.73
0.663
SVM
1.0
0.5
Linear SVM
0.522
0.512
Bayesian
0.514
0.512
Decision Tree
1.0
0.543
Random Forest
1.0
0.743
The table clearly indicates that SVM and naïve
Bayesian approaches are not performing well on the
data. Decision tree is strongly overfit and useless on
Died in 30 days Not died in 30 days
N = 1425 N = 20226
Cardiac dysrhythmias
42.25% 36.73% 1.26
Acute and unspecified renal failure
37.05% 21.12% 2.20
Essential hypertension
39.16% 52.57% 0.58
Respiratory failure; insufficiency; arrest (adult)
33.40% 17.88% 2.30
Congestive heart failure; nonhypertensive
22.60% 16.28% 1.50
Pneumonia (except that caused by TB or STD)
25.40% 12.66% 2.35
Urinary tract infections
24.70% 16.20% 1.70
COPD
24.84% 17.75% 1.53
Diabetes mellitus without complication
25.47% 24.55% 1.05
Deficiency and other anemia
29.19% 22.87% 1.39
Fluid and electrolyte disorders
27.93% 20.52% 1.50
Disorders of lipid metabolism
26.95% 39.20% 0.57
Coronary atherosclerosis and other heart disease 18.67% 23.09% 0.76
Comorbidities
LR
C-Lace: Computational Model to Predict 30-Day Post-Hospitalization Mortality
171

testing data. Logistic regression preforms reasonably
on both sets. Although its performance on testing data
is below desired level.
Random Forest (Breiman, 2001) has consistently
shown the highest accuracy on testing data, despite
clear overfit. Detailed analysis of the model presented
in Section 4 shows that the model is stable and
appropriate. Based on the result, the remainder of this
paper will focus on using Random Forest as the
prediction algorithm. It is a well-studied approach,
previously used in healthcare (i.e., Gu et al., 2015), in
which large number of shallow decision trees are
generated based on subsets of data (both examples
and attributes). In our case, the best performance was
achieved when generating 1,000 trees.
3.2 Use of Administrative and Clinical
Data
The primary dataset used to test the research question
is MIMIC III (Johnson et al., 2016) which is part of
PhysioNet project (Goldberger et al., 2000). The
dataset includes a variety of patient and clinical
information about hospitalizations, ICU, and patient
history. MIMIC III comprises over 58,000 hospital
admissions for 38,645 adults and 7,875 neonates. The
data spans June 2001 - October 2012. The rationale
of using MIMIC III in this project is that it includes
much more complex and diverse information than
typically found in claims data. One of our goals is to
illustrate that learning models from such data using
the described method leads to better results than those
that can be obtained from claims only data.
In the second set of experiments we tested if
addition of clinical data (lab values) to administrative
data (coded diagnoses) improves accuracy of
prediction of 30-day mortality. Inclusion of lab
values is consistent with existing models such as
APACHE II.
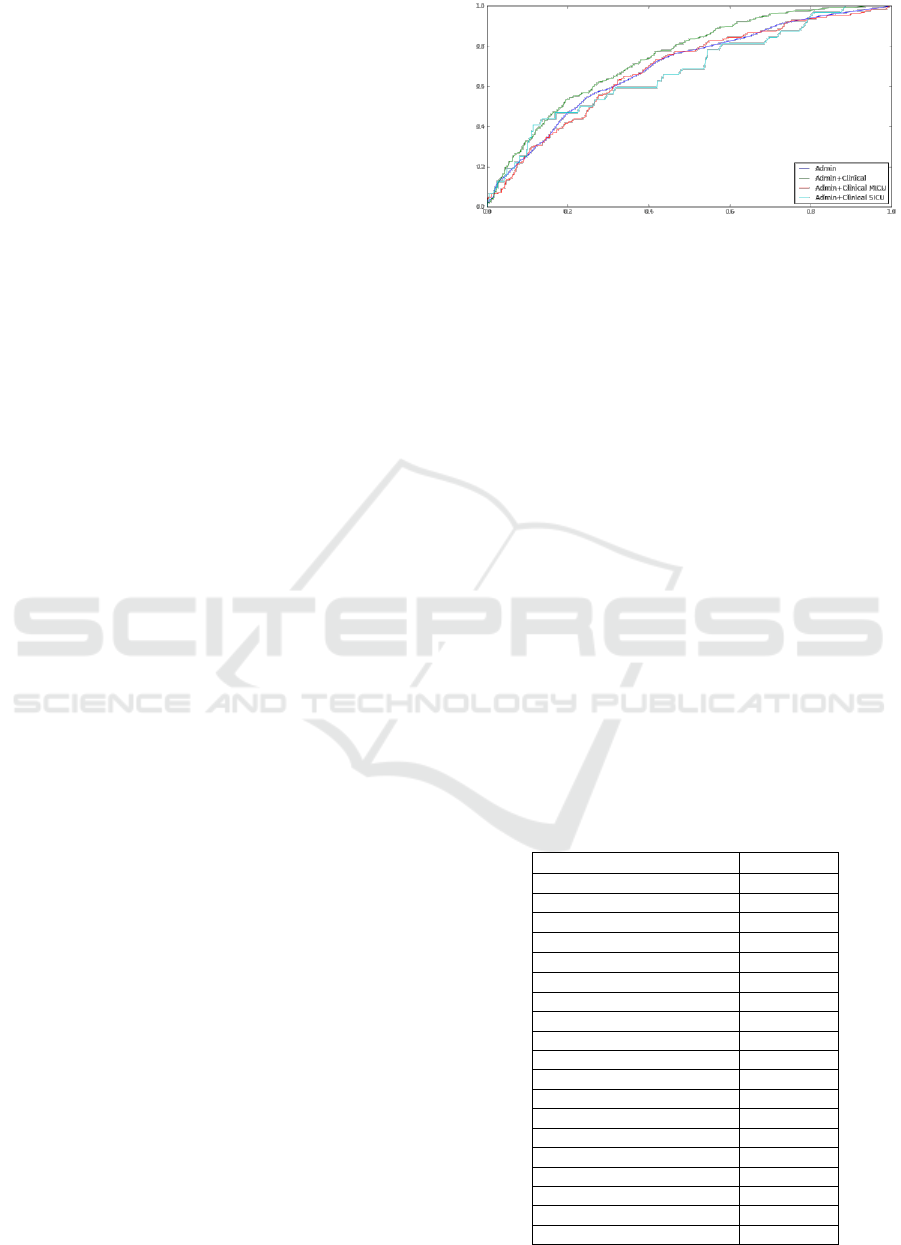
The results indicate that addition of clinical data
makes small difference in the accuracy. The AUC
increases from 0.72 to 0.74. The ROC for combined
administrative and clinical data is consistently above
one for administrative data only, as shown in Figure
1. Interestingly, when applied to Medical Intensive
Care Unit (MICU) and Surgical Intensive Care Unit
(SICU) patients only, the accuracy worsens. While
contradictory to the fact that these are two distinct
types of patients and separate modeling should
improve accuracy, this discrepancy can be explained
by the amount of data available and thus overfitting
of models.
Figure 1: Receiver-operator curves for four variants of C-
LACE model learned from administrative data only and
administrative and clinical data. Curves for MICU and
SICU patients are additionally presented.
3.3 Minimum C-LACE Model
Finally, we investigated possibility of reducing
number of attributes needed to accurately predict 30-
day mortality. Such a reduction is important for
simplification of the model and, as described in
Section 4, allows for creation of online calculator in
which data can be entered manually.
All 308 attributes used in the full model were
ranked based on their Mean Decrease Impurity
calculated by the Random Forest model. It is a
standard measure reported by RF after forests are
built. We created a set of models while increasing
number of attributes until the accuracy became
comparable to one in full model. This resulted in
selection of top 20 attributes listed in Table 3 along
with their weights. The table also includes counts of
patients and likelihood ratio as additional measure of
attribute quality.
Table 3: Selected top 20 attributes along with their
importance.
Feature
Importance
Age
0.0452
HOSPITAL_LOS
0.0346
MICU_LOS
0.0320
CCU_LOS
0.0177
CCS 106
0.0176
CCS 157
0.0169
CCS 98
0.0159
ADMISSION_LOCATION
0.0157
CCS 131
0.0152
CCS 108
0.0145
CCS 122
0.0133
SICU_LOS
0.0130
CCS 159
0.0129
CCS 127
0.0127
CCS 49
0.0127
CSRU_LOS
0.0126
CCS 59
0.0123
CCS 55
0.0123
CCS 53
0.0110
HEALTHINF 2017 - 10th International Conference on Health Informatics
172
The AUC of the model based only on age was
0.516 which is basically a random guess based on
prior class distribution. Similarly, the AUC of the
model based on Age and Length of Hospital Stay was
0.576. Interestingly models based on 5 and 10 top
attributes performed very close to each other with
AUC values of 0.6961 and 0.697, respectively.
Finally, the model based on 20 attributes performed
only slightly worse than one based on all 308
attributes (AUCs 0.734 and 0.743 respectively).
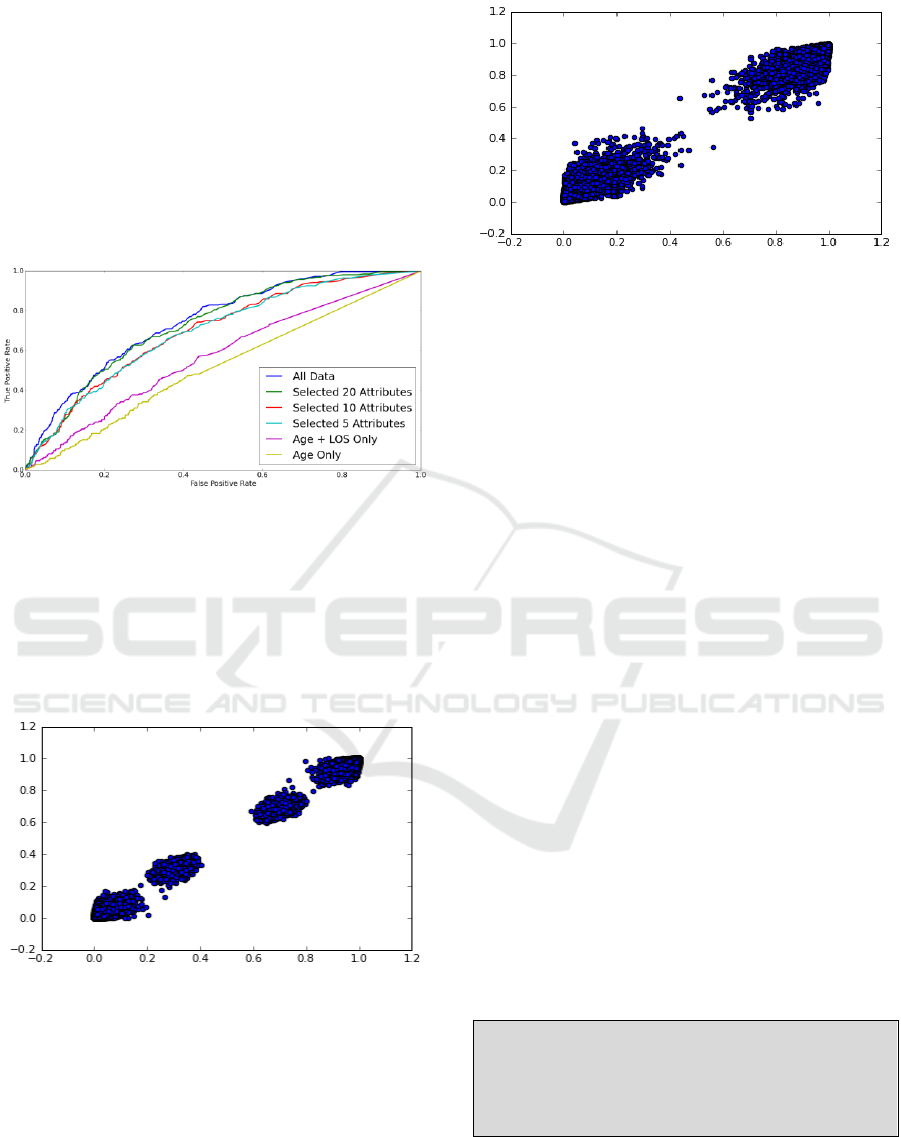
Figure 2 below illustrates ROC for these models.
Figure 2: Accuracy of models for different selection of
attributes given as ROC.
Additional analysis indicates that in fact predicted
probabilities from both models are very close. When
applied to training data Mean Squared Error (MSE)
between probabilities of 30-day mortality calculated
between both models was 0.000439 as illustrated in
scatterplot in Figure 3.
Figure 3: Comparison of probabilities of C-LACE and
Minimum C-LACE on training data.
Similarly, when compared on testing data the
MSE between the two models was 0.00335 as shown
in Figure 4. While there is a slight difference in the
predicted probabilities, the data are clearly clustered
into two groups that correspond to low and high risks
of mortality. Assignment to these groups is virtually
identical regardless of models used.
Figure 4: Comparison of probabilities of C-LACE and
Minimum C-LACE on testing data.
The above analysis indicates that the two models
are almost identical in terms of predictions, thus the
simpler of the models should be used.
4 MODEL ANALYSIS
In addition to standard testing of the created model
presented in the previous section, this section
discusses a more detailed analysis of the created
Minimum C-LACE model. The goal is to understand
the model’s behavior and its sensitivity to changes in
input attributes.
The first set of experiments was to investigate
how probabilities of 30-day mortality depend on
changes in single variables. This is particularly
important for continuous variables for which model
should be “smooth” and not produce sudden changes
in output probabilities. This property can be
investigated be applying the model to large simulated
data and comparing output to distribution of values in
real dataset.
First created simulated dataset was completely
random, that is, each input attribute was assigned
uniformly a random value from list of allowed values
for that attribute with exception for one attribute
being controlled. For example, generation of
simulated data to test age attribute followed the
procedure:
for a = min(age) to max(age):
Generate 1,000 random examples:
age = a
for each attribute x other than age:
x = random(domain(x))
After simulated dataset is generated, C-LACE
model is applied to predict mortality probabilities.
These probabilities can then be investigated to check
model’s behavior based on changes in age.
C-Lace: Computational Model to Predict 30-Day Post-Hospitalization Mortality
173
Obviously, accuracy measures are not applicable to
this simulated data since no true answer is known.
The result is shown in Figure 5, which also includes
distribution of average values depending on age in
training, testing and complete data.
One can immediately note that the probabilities
based on “completely random” simulated data are
much higher than those in real data. This is correct,
because a completely randomly generated patient is
much “sicker” than real patients due to the way data
are generated. The data on the plot shows that the
model is smooth in regard to changes of probability
with age. An interesting fact about model is that
probabilities are somewhat higher for the lowest
allowed value of age, namely 65.
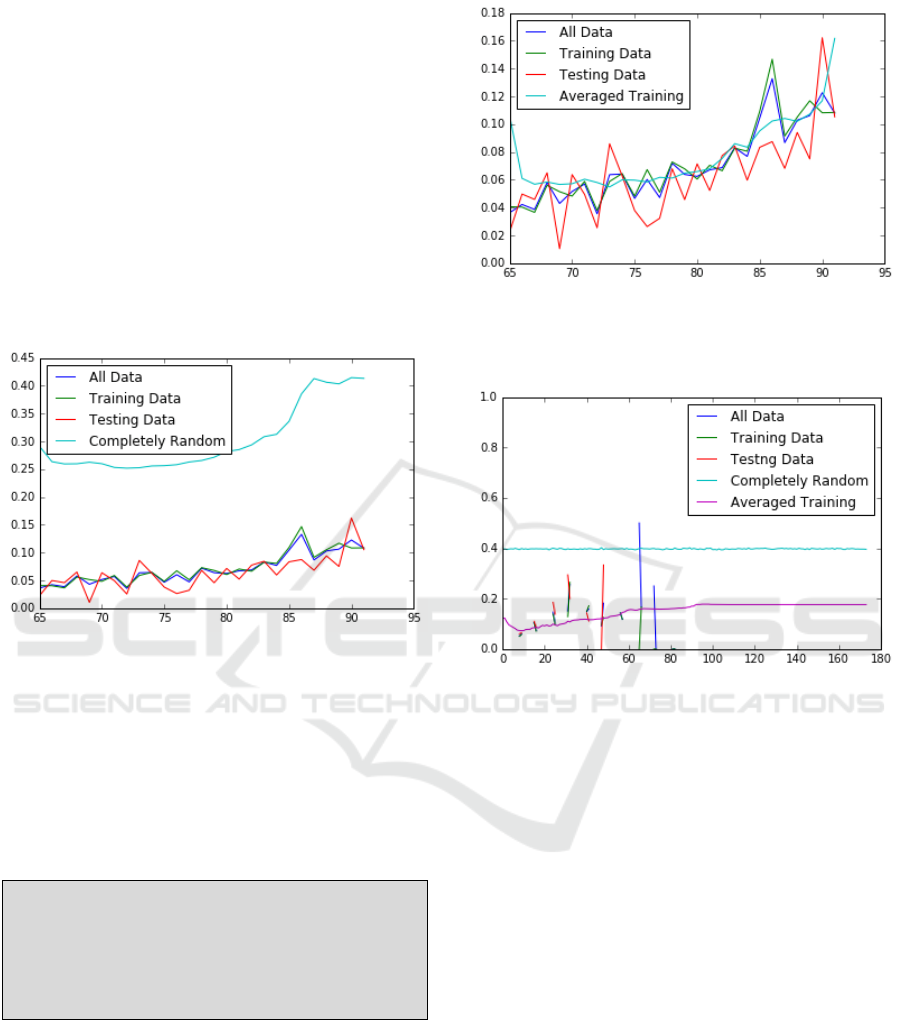
Figure 5: Distribution of predicted probability of 30-day
mortality based on patient age for completely random
simulated data compared with real data.
The second (averaged training) method used to
generate simulated data started with original dataset
used for training C-LACE model and multiplied the
data by copying all examples for each fixed age and
applying low probability random distortion to all
other attributes.
for a = min(age) to max(age):
For each example in training data:
Copy the example
age = a
for each attribute x other than age:
distort x
One can notice that probabilities of mortality in
the simulated data are no longer higher than those of
real data. This is due to the fact that all attributes other
than age are distributed as in the original dataset
(Figure 6). In the plot, one can immediately see that
there is a similar “jump” of probability at the age of
65 indicating possible instability of model there.
Figure 6: Distribution of predicted probability of 30-day
mortality based on patient age for averaged training
simulated data compared with real data.
Figure 7: Distribution of predicted probability of 30-day
mortality based on hospital length of stay for completely
random simulated data compared with averaged training
data and actual data.
The same methodology for creating completely
random and averaged training simulated data has
been applied to other attributes in the data with
similar results. One interesting result was obtained
when simulating data for fixed hospital length of stay
(LOS) shown in Figure 7. When applied to
completely random data, LOS has absolutely no
effect on predicted probability (straight line on the
plot). Interestingly, on simulated averaged training
data, LOS shows clear trend. One possible
explanation of this fact is that within the model LOS
is strongly confounded with other attributes. The
visible trend is in fact one of other attributes
interacting with LOS to affect predicted mortality
indirectly. Finally, a number of colored randomly
looking lines in Figure 7 show that in the original data
there is no clear pattern of how LOS affects predicted
30-day mortality.
The fact that when working with simulated data
probabilities output by the model are smooth,
HEALTHINF 2017 - 10th International Conference on Health Informatics
174
confirms the hypothesis that the constructed C-LACE
model is stable.
4.1 Analysis of Errors
An interesting and important question concerns
finding cases for which the model makes mistakes. If
successful, such analysis may allow for predicting
when C-LACE is more likely to make a mistake, and
thus preventing it.
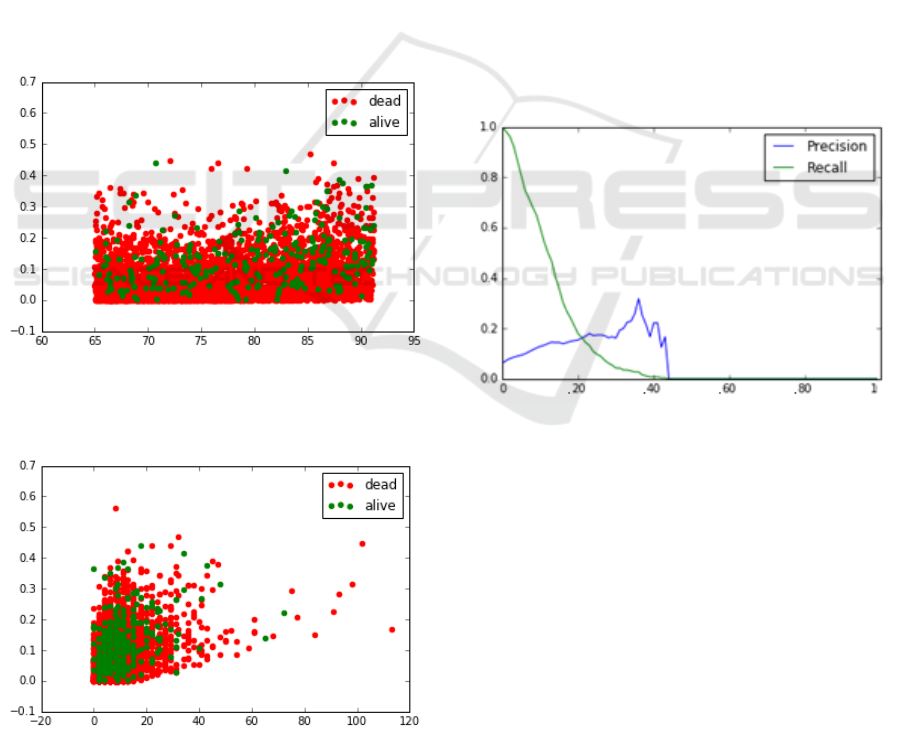
As shown in Figures 8 and 9, there is basically no
pattern on when the model makes mistakes based on
distribution of age and length of hospital stay. In both
figures, green dots representing patients who died
should be clustered towards the top, and red ones
representing alive patients towards the bottom. The
distribution errors in the model (how far green dots
are from the top) is practically uniform with respect
to age. While the distribution of hospital length of
stay is clearly positively skewed, there seems to be no
pattern in when errors are made (Figure 9).
Figure 8: Predicted probabilities of 30-day mortality for
training data in relation to patient age. Color of dots
represents true class.
Figure 9: Predicted probabilities of 30-day mortality for
training data in relation to hospital length of stay. Color of
dots represents true class.
Secondary model was learned from data to predict
when C-LACE is likely to misclassify positive
mortality examples. Specifically, it was built from
data labeled as correct classification/misclassification
of testing data used to evaluate C-LACE. The
secondary model has been learned using logistic
regression. Following the standard procedure the
misclassification data was split into training (80%)
and testing (20%). When tested, the model showed
very high promise of predicting when C-LACE is
likely to make mistakes. It achieved AUC 0.867 on
misclassification training and AUC 0.858 on
misclassification testing data.
The final set of performed tests investigated
optimal classification threshold based on precision
and recall. Using C-LACE it is possible to achieve
any value or recall, precision in general stays very
low as shown in Figure 10. The figure indicates that
selection of classification threshold for C-LACE
around 0.1 may be the most reasonable. More detailed
cost-benefit analysis of false positives and false
negatives of the model is needed to arrive at final
threshold applicable for final use.
Figure 10: Analysis of Precision and Recall of the
Minimum C-LACE model on testing data.
5 ONLINE CALCULATOR
In order for other researchers to test the developed
mortality prediction models, an online calculator
which includes Minimum and Full C-LACE models
was created. The minimum model is available
through a web form that can be used by entering data,
as well as Application Programing Interface (API) for
automated use. The full model is available only
through an API, since it is unlikely for anyone to
answer 308 questions on a web form. At this stage,
the online calculator is intended only for research
purposes and not for clinical use, since additional
C-Lace: Computational Model to Predict 30-Day Post-Hospitalization Mortality
175

validation is needed. The online calculator is
available at the website http://hi.gmu.edu/cgi-
bin/calculatros/c-lace/c-lace.cgi.
Figure 11: Design of the simple form used to enter patient
and hospitalization information.
Simple online form (Figure 11) is used to enter
patient and hospitalization characteristics. The entry
is split into sections related to length of stay in
hospital and specific ICUs, age, admission location
and selected conditions most predictive of 30-day
mortality. After submitting the form, user is provided
with estimated probability of 30-day mortality.
Because of the way the data was analyzed, the
calculator is intended to be used at the time of hospital
discharge.
It is important to note that within the scope of this
project it was impossible to completely test the
calculator and in particular assess its impact on
patient care. Thus, the site contains a disclaimer that
the calculator is intended to be used only for research
purposes.
6 CONCLUSIONS
This paper presented construction and analysis of C-
LACE method for predicting probability of 30-day
post-hospitalization mortality. The presented solution
based on application of Random Forest algorithm
gives accuracy comparable to other methods
available in the literature and superior to accuracy of
the original LACE index. It shows that Minimum C-
LACE, a 20-attributes version of the presented
method, achieves the same results as one that uses
308 attributes.
Detailed analysis of the constructed model shows
that the model is not sensitive to changes in values of
key variables and, in fact, smoothens the data (the
most visible for length of stay). While the accuracy of
the model precludes its use completely
independently, it is a reasonable improvement over
popular LACE method. The model can be used to
inform clinicians when performing patient risk
assessment. Analysis has indicated that it may be
possible to automatically assess classification errors
from the model, though additional work is needed in
this area.
The current continuation of research proceeds in
two main directions:
- Possible improvement of the model accuracy by
using additional clinical variables. There is
significant work that remains to be done in the area
of incorporating detailed clinical information and
patient notes with specific focus on temporal aspect
of the data. In the presented Minimum C-LACE
model, no clinical attributes were included, which
may be result of oversimplification of how the
values were coded (see Section 2.2).
- Analysis of how the model should be presented to
end-users so they understand predicted
probabilities and model limitations. The latter is
particularly important to make the presented online
calculator useful.
ACKNOWLEDGEMENTS
This project has been supported by the LMI-
Academic partnership program grant. The authors
thank Donna Norfleet, Chris Bistline, Brent Auble
and all those who participated in our monthly
meetings for their support and feedback that helped
improve the project.
The authors thank anonymous reviewers whose
feedback helped improve the paper, particularly by
pointing to relevant previously published literature.
REFERENCES
AHRQ (2016) https://www.hcup-us.ahrq.gov/tools
software/ccs/ccs.jsp
Au, AG, McAlister, FA, Bakal, JA, et al. 2012. Predicting
the Risk of Unplanned Readmission or Death within 30
Days of Discharge After a Heart Failure
Hospitalization. American Heart Journal. 164(3):365-
372.
Breiman, L., 2001. Random forests. Machine learning,
45(1), 5-32.
Cooper, Gregory F., et al., 1997. An evaluation of machine-
learning methods for predicting pneumonia mortality.
Artificial intelligence in medicine 9.2. 107-138.
Davis, RB, Iezzoni LI, Phillips RS, Reiley P, et al. 1995.
Predicting in-hospital mortality: the importance of
functional status information. Med Care.33:906-921.
Goldberger AL, et al., 2000. PhysioBank, PhysioToolkit,
and PhysioNet: Components of a New Research
Resource for Complex Physiologic
HEALTHINF 2017 - 10th International Conference on Health Informatics
176

Signals. Circulation 101(23):e215-e220 [Circulation
Electronic
Pages; http://circ.ahajournals.org/content/101/23/e215.
full
Gu, W., Vieira, A. R., Hoekstra, R. M., Griffin, P. M., &
Cole, D. , 2015. Use of random forest to estimate
population attributable fractions from a case-control
study of Salmonella enterica serotype Enteritidis
infections. Epidemiology and Infection, 1-9.
Ho KM, Dobb GJ, Knuiman M, et al. 2006. A comparison
of admission and worst 24-hour Acute Physiology and
Chronic Health Evaluation II scores
in predicting hospital mortality: a retrospective cohort
study. Crit Care; 10:R4.
Iezzoni LI. Risk Adjustment for Measuring Health
Outcomes Ann Arbor. 1994. Mich: Health
Administration Press.
Inouye, S.K., Peduzzi, P., Robison, J., Hughes, J., Horwitz,
R., Concato, J, MPH, 1998. Importance of Functional
Measures in Predicting Mortality Among Older
Hospitalized Patients. JAMA. 1998;279(15):1187-
1193. doi:10.1001/jama.279.15.1187.
Johnson AEW, Pollard TJ, Shen L, Lehman L, Feng M,
Ghassemi M, Moody B, Szolovits P, Celi LA, and Mark
RG. 2016. MIMIC-III, a freely accessible critical care
database. Scientific Data. 10.1038/sdata.2016.35.
Kuzniewicz MW1, Vasilevskis EE, Lane R, Dean
ML, Trivedi NG, Rennie DJ, Clay T, Kotler
PL, Dudley RA. 2008. Variation in ICU risk-adjusted
mortality: impact of methods of assessment and
potential confounders. Chest. 2008;133(6):1319-27.
doi: 10.1378/chest.07-3061.
Levy, C., Kheirbek, R,, Alemi, F., Wojtusiak, J., Sutton, B,,
Williams, A.R. and Williams, A., 2015. Predictors of
six-month mortality among nursing home residents:
diagnoses may be more predictive than functional
disability. Journal of Palliative Medicine, 18(2), 100-6.
Moreno, RP, Metnitz, P.G., Almeida, E, et al., 2005. SAPS
3--From evaluation of the patient to evaluation of the
intensive care unit. Part 2: Development of a prognostic
model for hospital mortality at ICU admission.
Intensive Care Med; 31:1345.
Moskovitch, R. and Shahar, Y., 2015. Classification-driven
temporal discretization of multivariate time series. Data
Mining and Knowledge Discovery, 29(4), pp.871-913.
Ngufor, C., Wojtusiak, J., Hooker, A., Oz, T. and Hadley,
J., 2014. Extreme Logistic Regression: A Large Scale
Learning Algorithm with Application to Prostate
Cancer Mortality Prediction. Proceedings of the 27th
International Florida Artificial Intelligence Research
Society Conference.
Rocker, G., Cook, D., Sjokvist, V., Weaver, B., Finfer, S.,
McDonald, E., Marshall, J., Kirby, A., Levy, M., at al.,
2004. Clinician Predictions of Intensive Care Unit
Mortality, Crit Care Med. 2004;32 (5)1
Rose, Sherri, 2013. "Mortality risk score prediction in an
elderly population using machine learning." American
journal of epidemiology 177.5. 443-452.
Taylor, R. Andrew, et al. 2016. Prediction of In‐ hospital
Mortality in Emergency Department Patients with
Sepsis: A Local Big Data–Driven, Machine Learning
Approach. Academic Emergency Medicine.
Van Walraven, C., Dhalla, IA, Bell, C., et al., 2010.
Derivation and validation of an index to predict early
death or unplanned readmission after discharge from
hospital to the community. CMAJ 2010 Apr 6; 182(6):
551–557. doi: 10.1503/cmaj.091117. PMCID:
PMC2845681
Van Walraven C, Wong J, Forster AJ. 2012. LACE+
index: extension of a validated index to predict early
death or urgent readmission after hospital discharge
using administrative data. Open Med. 2012 Jul
19;6(3):e80-90. PMCID: PMC3659212
Vasilevskis EE, Kuzniewicz MW, Cason BA, Lane
RK, Dean ML, Clay T, Rennie DJ, Vittinghoff
E, Dudley RA. 2009. Mortality probability model III
and simplified acute physiology score II: assessing their
value in predicting length of stay and comparison to
APACHE IV. Chest. 136(1):89-101. doi:
10.1378/chest.08-2591.
Verduijn, M., Sacchi, L., Peek, N., Bellazzi, R., de Jonge,
E. and de Mol, B.A., 2007. Temporal abstraction for
feature extraction: A comparative case study in
prediction from intensive care monitoring data.
Artificial intelligence in medicine, 41(1), pp.1-12.
Wojtusiak, J., Elashkar, E., Mogharab, R., 2016. Integrating
Complex Health Data for Analytics. The Machine
Learning and Inference Laboratory, electronic
circulation. MLI 16-1.
C-Lace: Computational Model to Predict 30-Day Post-Hospitalization Mortality
177
