
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
I. Fayerverker and T. Mor
Department of Computer Science, Technion, 3200003, Haifa, Israel
Keywords:
Autocatalytic Sets, Artificial Life, Origin of Life, Universal Replicator, Genetic Code, Evolution, Origin of
Complexity, Translation.
Abstract:
The genetic code and genetic evolution are at the core of complexity in biology, however, there is no reasonable
explanation yet for the emergence of the genetic code. We present here a possible scenario accounting for the
emergence of “coded life” in nature: We describe the emergence of the genetic code from molecular evolution
(prior to genetic evolution). This process is based on increase in concentration of chemical self-replicating sets
of molecules, located within (probably non-biological) compartments. Our scenario is obtained by combining
the conceptual idea of “code-prompting autocatalytic sets” (Agmon and Mor, 2015), with recent results about
non-enzymatic template replication methods (Prywes et al, 2016), possibly relevant to the prebiotic stage
preceding RNA-world. In the scenario described here, we often use computer science viewpoint and abstraction:
We consider sets of strings composed of letters, such that each letter represents a molecular building block
— mainly nucleotides and amino acids, and each string represents a more complex molecule which is some
concatenation of the simpler molecules represented by letters; the biochemical rules are described in an abstract
language of rules and statistics of letters and strings. We then suggest a novel path, containing several phases,
for the emergence of “coded life”.
1 INTRODUCTION
A major objective of scientific endeavor is the elu-
cidation of the origin of life on Earth (Schroedinger,
1944; Woese, 1967; Orgel, 1968; Dyson, 1985; Gilbert,
1986; Koshland, 2002). There is still no standard defi-
nition of the term “Life’ ’(Schroedinger, 1944; Dyson,
1985; Koshland, 2002), and there is no “standard
model” of the origin of life. There is however a rather
general agreement that RNA preceded DNA (Gilbert,
1986; Lazcano et al., 1988; Horning and Joyce, 2016),
and that “it all started” from a prebiotic primordial
assembly of molecules (also known as ”the primor-
dial soup” or ”the prebiotic soup”). It is also believed
that evolution occurred first in populations of complex
molecules (Vasas et al., 2012), and potentially in non-
biological compartments (Koonin and Martin, 2005),
and only later in “cellular proto-organisms”.
Among the pillars of “life” one can surely list
compartmentalization, replication, evolution, muta-
tions, a code, energy consumption, and active trans-
port. Among the methods that enable thinking and
analyzing models for defining life and/or for the emer-
gence of life one can surely list continuation of evo-
lution (with cases of jumps as well) and search for
marks left in contemporary bio-molecules (such as
the ribosome and the polymerase) that are common
to all living organisms. A work by (Agmon and Mor,
2015) added recently the method of abstraction, and
suggested a model dealing with the first five pillars
mentioned above
1
. We suggest here an improvement
over (Agmon and Mor, 2015) based on a new experi-
mental method for template replication suggested and
implemented by (Prywes et al., 2016).
When and how genetically-coded proto-organisms,
which we call here “coded life”, first appeared along
the path of evolution is still not clear. Although var-
ious different models concerned with the emergence
of life, e.g. (Woese, 1967; Orgel, 1968; Crick, 1968;
Gilbert, 1986; Kunin, 2000; Segr
´
e et al., 2001; Koonin
and Martin, 2005; Ikehara, 2005; Yarus et al., 2005;
Koonin and Novozhilov, 2009; van der Gulik et al.,
2009; Kauffman, 2011; Vasas et al., 2012), have re-
sulted in significant progress over the last decades
in clarifying alternatives regarding the origin of life,
none of the models presents a complete scenario for
the emergence of life. In particular, the emergence of
the genetic code (Woese, 1967; Orgel, 1968; Crick,
1
For an interesting work dealing thoroughly with the last
two pillars while also clarifying a potential path from non-
biological compartments to biological ones see (Lane and
Martin, 2012; Lane, 2015).
Fayerverker, I. and Mor, T.
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life.
DOI: 10.5220/0006681300530063
In Proceedings of the 3rd International Conference on Complexity, Future Information Systems and Risk (COMPLEXIS 2018), pages 53-63
ISBN: 978-989-758-297-4
Copyright
c
2019 by SCITEPRESS – Science and Technology Publications, Lda. All rights reserved
53

1968) remains a major open question (see for exam-
ple (Kunin, 2000; Lahav et al., 2001; Yarus et al.,
2005; Koonin and Novozhilov, 2009; Rouch, 2014)).
Although there are various models regarding very early
stages of the emergence of life in which there are no
peptides involved, such as RNA-world (Gilbert, 1986;
Lazcano et al., 1988; Horning and Joyce, 2016) and
lipid-world (Segr
´
e et al., 2001), it seems natural that
the world during the emergence of translation and of
the genetic code must have had at least two types of
highly-relevant letters (molecules) — amino acids and
nucleotides, and strings formed from these basic build-
ing blocks (Lahav et al., 2001; Agmon and Mor, 2015).
It is well known that some sets of strings together
with their reactions form autocatalytic sets (Kauffman,
1986; Hordijk and Steel, 2004; Hordijk et al., 2010;
Hordijk et al., 2011; Kauffman, 2011; Vasas et al.,
2012). Note that we use here the abbreviation ACS to
describe autocatalytic set (singular), and autocatalytic
sets (in plural).
Following (Agmon and Mor, 2015), we find sev-
eral unique autocatalytic sets of strings comprised of
these two types of letters, amino acids and nucleotides,
that prompt the emergence of a genetic code. Agmon
and Mor’s model seems to be connected to contem-
porary biology and “life as we know it”, yet it is less
connected to the chemical era (i.e., to RNA world or
to pre-RNA-world). Their work presented an ACS
of strings (molecules) which is a probable possibility
to be the base of the contemporary genetic code; it
is named “Code-Prompting-ACS”, or COPACS. This
set is unique since it is the only current model that
describes the emergence of the genetic code in detail.
However, their model does not present a clear evo-
lutionary path from the simplest molecular evolution
to COPACS: In their model, two rare events had to
happen, namely two long molecules need to randomly
and spontaneously be generated from the primordial
soup, the proto-ribosome (
R
0
in that paper, and here)
and the proto-polymerase (
P
0
in that paper, and here)
or more precisely, its coding in a messenger RNA. The
two molecules are highly complex since (a point not
clearly specified in (Agmon and Mor, 2015)) both need
to be motoric:
R
0
need to be motoric in order to move
onto the messenger RNA during translation, and
P
0
need to be motoric in order to move on the template
during template replication.
The model we present here closes these gaps and
suggests a reasonable link of (Agmon and Mor, 2015)
COPACS to the chemical era, relying only on a sin-
gle rare event; here we show that only a single highly
unlikely molecule had to appear at random from the
primordial soup — the motoric
R
0
. Therefore, the
COPACS presented here suggest a relatively easy and
clear path (even if highly hypothetical for now) of
continuous evolution from the chemical era to coded
life and hence contribute an important phase to the
comprehension of the emergence of life as we know it.
Also, while Agmon and Mor suggested that the first
code word must be the messenger RNA coding the
polymerase, our COPACS are much more flexible and
open various options for the first code word — it may
be (the coding of) any one of various peptides that sig-
nificantly improves the catalysis of (RNA-catalyzed)
template replication, as is explained in details in Phase
3, of Section 5.
The content of the rest of this paper is as follows:
In Section 2 we discuss Kauffman’s ACS, and the
method of Agmon and Mor for code prompting ACS.
In Section 3 we define the notation and rules of ”letters
and strings” model for ”digital abstraction”, used in
this paper, as well as in (Agmon and Mor, 2015)
. In Section 4 we discuss the method of (Prywes
et al., 2016) for non-enzymatic template replication.
In Section 5 we present our scenario for the evolution
of the genetic code, phase by phase. In Section 6 we
discuss our results, and potential future research.
2 ACS AND COPACS
Originally two methods, template replication (Watson
and Crick, 1953) and autocatalytic sets (ACS) (Kauff-
man, 1986; Hordijk and Steel, 2004; Hordijk et al.,
2010; Hordijk et al., 2011; Kauffman, 2011; Vasas
et al., 2012), were presented as competitive mod-
els for basic evolution from the prebiotic soup, into
a much richer organic prebiotic environment [Note
that (Hordijk et al., 2010; Hordijk et al., 2011) dis-
cuss the template replication as well]. ACS was first
suggested as a model for replication of peptides (Kauff-
man, 1986). However, around the same time, ri-
bozymes, i.e., RNA molecules that act as enzymes
were found (Guerrier-Takada et al., 1983). As a result,
template replication became a leading and fully agreed
method for basic evolution. And on the other hand,
ACS of RNA strings was eventually also considered
in later work on ACS. Additionally, variants of both
models (e.g., (Kunin, 2000; Lahav et al., 2001; Rouch,
2014; Agmon and Mor, 2015)) explored the possibility
of a world in which RNA and small peptides evolved
together.
ACS here means a
complete
catalytic set of
molecules and reactions, where no outside help (in
terms of the required molecules) is needed for the repli-
cation process [for more formal details see: (Hordijk
et al., 2010; Hordijk et al., 2011; Vasas et al., 2012)].
Intuitively speaking, the set of molecules of an
COMPLEXIS 2018 - 3rd International Conference on Complexity, Future Information Systems and Risk
54
ACS includes some given food-molecules (available
in large quantities), and in addition, some non-food
molecules, all of which are generated, directly or as
a result of series of reactions, from the given set of
food-molecules. In the general model of catalysis used
by (Kauffman, 1986; Hordijk et al., 2010; Hordijk
et al., 2011), a reaction is either catalyzed or not
catalyzed by a given molecule: For simplicity, non-
catalyzed processes are commonly excluded from the
set (Kauffman, 1986) since catalysis enhances the rate
of a reaction by several orders of magnitude, thus es-
sentially abolishing the opposite process ( i.e. from
the product to the reactants). In addition, a vital point
in defining any ACS is that each reaction in the ACS
must be catalyzed by at least one molecule in the ACS.
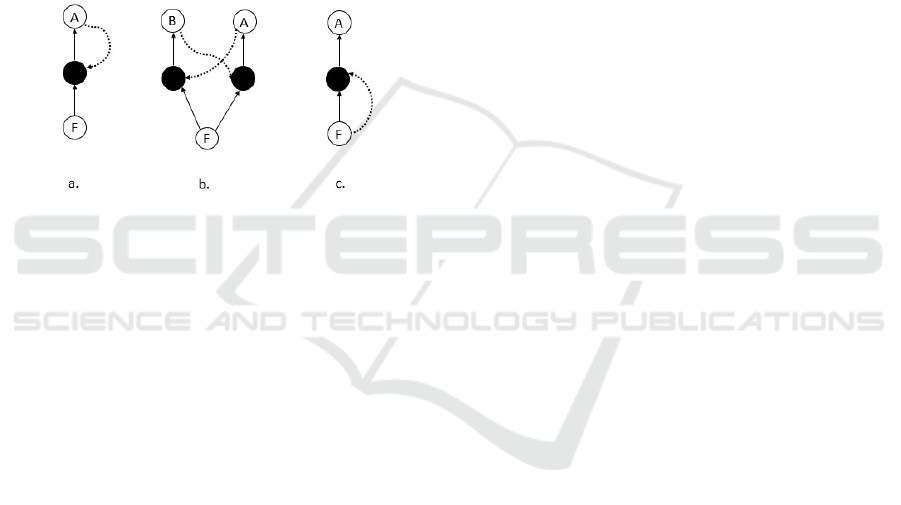
(See Figure 1).
Figure 1: Simple examples of ACS. Following (Hordijk et al.,
2010; Hordijk et al., 2011; Vasas et al., 2012) we present
here several examples. Black dots are used here for reactions,
empty circles for molecules or sets of molecules (e.g.
F
for
the set of all food molecules). Full lines indicate input and
output of a reaction, and dotted lines – catalytic processes.
a. A single autocatalytic molecule
A
catalyzing its own
formation. b. Molecules
A
and
B
catalyze the formation
of each other. This is the simplest non-trivial ACS — a
two-member autocatalytic loop. c. A single autocatalytic set
generating molecule
A
while the process is catalyzed by one
of the members of the food set.
Of course a more careful analysis of catalysis by
splitting into stronger and weaker strengths of catalysis
is also possible, however commonly is not required.
For our purpose here, in just one case — the template
replication defined in Phase 3, in Section 5, we do
need such a split of the catalysis into weak catalysis
and strong catalysis.
3 THE LETTERS AND STRINGS
MODEL
In this section we follow (Agmon and Mor, 2015)
and and shortly describe their offered notation and
rules of ”letters and strings” model for ”digital abstrac-
tion”. As is common for describing complex systems
in computer science and information theory, we use an
abstraction — instead of looking deep into the physics,
chemistry, and biochemistry involved, we treat the ori-
gin of life as (biochemically-motivated) statistics and
rules regarding letters and strings. This model may be
viewed as a “digital abstraction” of the biochemistry
involved in the processes we describe here.
In general, “abstraction” is a method of treating
complex systems at several different levels, in our case,
a physical level, the chemistry level, the digital level
discussed here, and then the genetic code level. The
higher level is always used to simplify things, when
describing a highly complex system, while the lower
level is used to clarify where the rules and definitions
of the higher level came from. In our “digital level”,
monomers are letters, polymers are strings of letters,
and reactions are usually simplified to binary opera-
tions (yes/no).
3.1 Letters and Strings Inside
Compartments
The most important players in our model are strings
built from two types of letters (namely two types of
molecules),
r
for RNA nucleotides, and
p
for amino
acids. As in the ACS model (Kauffman, 1986), as
well as in other models (Koonin and Martin, 2005),
we assume these molecules are located within a non-
biological compartment; for the steps of emergence
described here, the properties of the compartment are
not highly important, as long as sometimes compart-
ments are generated around some portion of the pre-
biotic soup of molecules and sometimes they are de-
stroyed/dissolved. We assume a small number of dif-
ferent letters: there are four types of
r
letters and at
most ten types of p letters.
3.2 Letters and Strings — Their
Characteristics
We define the following “digital” characteristics/rules
for these letters and strings:
1.
Both types of letters have the capability of being
combined into directed strings,
R
made of
r
let-
ters, and
P
consisting of
p
letters. Both types of
letters have directionality which can be described
as having a head and a tail, such that while form-
ing a string the head of one letter is connected to
the tail of a second letter of the same type. The
connection between neighboring letters along the
strings is named “backbone connection” for both
types of strings. These connections are assumed
to be strong, allowing the sustainability of the one-
dimensional string (1D).
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
55
Within each
R
string, each
r
letter can form back-
bone connections with any other
r
letter, such that
every arbitrary sequence of
r
’s is possible. Simi-
larly, every arbitrary sequence of
p
letters in each
P string is possible.
2.
We assume the existence of long random
R
strings
(e.g. couple of hundreds of
r
letters) in the environ-
ment (see (Ferris, 2002; Mast and Braun, 2010) for
the justification). Short
R
strings (say of length up
to 5) are highly common, while the probability of
specific longer strings becomes negligible as they
are longer, unless the specific string is catalyzed.
In contrast, the model we present here does not
rely on long P strings.
3.
In addition to their ability to be combined into
strings, the
R
and
P
strings can generate more com-
plex (2D and 3D) structures, by forming bonds
perpendicular to the string direction, namely per-
pendicular to the direction of the backbone connec-
tions. These connections, named “perpendicular
connections”, are assumed to be weaker than the
backbone connections.
(a)
Non-specific perpendicular connections (be-
tween two
p
letters or between two
r
letters)
exist, and these are the weakest connections.
(b)
In contrast to the strong backbone connections,
and to the weak non-specific perpendicular con-
nections, there is another type of “specific” per-
pendicular connections, only between specific
r
letters: Each letter
r
i
within a string can form
a perpendicular connection
only
with a single
“complementary letter” from the remaining set
of (three)
r
letters. Without loss of generality
we may assume here that
r
1
and
r
4
are com-
plementary to each other and that
r
2
and
r
3
are
complementary to each other. We will denote
the complementary nucleotide of given letter
r
by
r
0
. Thus, an
R
string has the potential to at-
tract specific
r
letters or another (specific) string,
to generate a ladder-like structure. If the at-
tracted string or formed string is precisely of the
same length as
R
, and it is the (letter by letter)
complementary string of R, we denote it as R
0
.
4.
There exists an attraction (called the stereochem-
ical attraction (Woese, 1965; Yarus et al., 2005;
Johnson and Wang, 2010)), between any
p
letter
and a specific triplet of
r
letters. Such a triplet of
r
letters is known as a “coding triplet”, and it is
specific per each letter of type p.
5.
A bond can form, between any specific
p
letter and
the last letter of a specific type of
R
string (that we
call
R
t
). The resulting string is called a “charged”
R
t
string — a
R
t
string with a
p
letter attached to
its end.
For more details and biochemical justifications for
this notation and set of rules - see (Agmon and Mor,
2015).
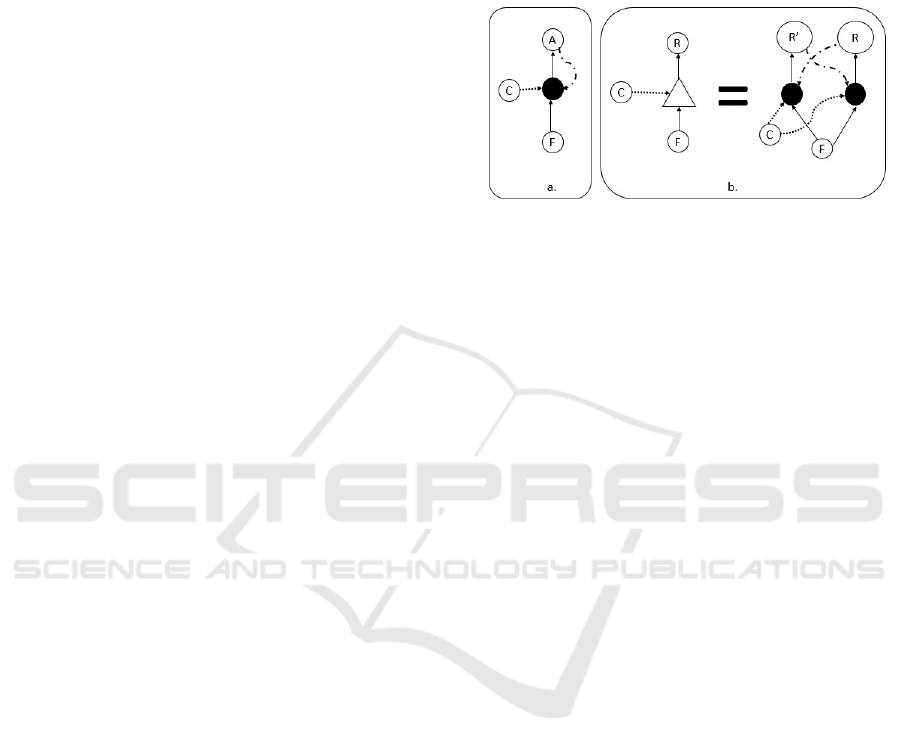
Figure 2: Template replication: a. We denote template repli-
cation using dash-dot-dash-dot line. Here, molecule
A
acts
as a template for its own replication while
C
is the catalyst
and
F
is the set of Food molecules. b. In the right side of
the equation we show template replication of
R
and
R
0
using
C
as a catalyst for both reactions and
F
as the set of food
molecules. To save us from complicated drawings whenever
template replication is shown, we define the triangle process
notation in the left side of the equation and use this notation
later on. Note that in both figures if
C
is in the food set or it
is one of the replicated molecules (
A
in a,
R
or it is
R
0
in b)
then the resulting set is an ACS, otherwise the resulting set
is not an ACS (since there is no catalytic process creates
C
).
4 TEMPLATE REPLICATION
WITH RNA CATALYSTS
In this section we look into template replication, going
beyond the digital abstraction, and discussing, still
rather briefly, the related biochemistry.
Many researchers assume RNA-world to exist prior
to today’s DNA-peptide world. Despite of differ-
ences in how various researchers precisely define the
RNA-world, the most important characteristic of RNA-
world is probably that RNA molecules that has 2D and
3D (relatively) complex structure acted as enzymes
when peptidic-enzymes did not yet exist. Such RNA-
enzymes are named ribozymes. For clarity, we refer
here to a peptide catalyzing template replication as the
proto-polymerase, and to an ribozyme catalyzing —
via an enzymatic process — the template replication,
as a replicase. In most models, as well as in current
biology, such enzymes are complex molecules, and
their performance is motoric; namely, after catalyzing
the addition of a single r letter - they move to the next
spot on the template.
It is believed by many researchers that having com-
plex ribozymes spontaneously emerging and then cat-
alyzing template replication might be highly improba-
ble, namely, their existence is implausible unless there
COMPLEXIS 2018 - 3rd International Conference on Complexity, Future Information Systems and Risk
56

is a previous step of template replication: Prior to an
RNA-world, or just in its beginning, it makes sense to
believe that short simple 1D RNA strings that has not
2D or 3D complex structure may act as catalysts. In
particular, such short 1D RNA strings may act as cata-
lysts for template replication, without being motoric.
4.1 The Replicase and the Trimers
When the RNA string that catalyzes template repli-
cation is the ribozyme replicase, we denote it as
R
∗
.
Alternatively, when an RNA string (or strings) catalyz-
ing template replication are simple 1D RNA strings,
that are then assumed to be part of the food molecules,
we shall not call them R
∗
; in relevant figures, but sim-
ply assume they are part of the Food molecules. If the
RNA string (or strings) catalyzing template replication
are simple 1D RNA strings, we do not have to assume
the existence of a world of sophisticated ribozymes,
and we can suggest how translation directly evolved
via simple molecular evolution.
4.2 Template Replication Via
Prywes-et-al. Extension
To support the belief that non-enzymatic template repli-
cation (namely, with no enzymes and no ribozymes),
existed in the chemistry era, some experiments tried to
check various non-enzymatic yet catalytic processes
for template replication. Let us focus here and provide
more details about one such non-enzymatic process
— the extension of an RNA string by a single letter
described by (Prywes et al., 2016).
Extension of the replicated RNA string may be
done step by step, by adding one
r
letter at each step or
by a process named ligation where longer RNA strings
might be added.
For simplicity, and also due to the results of Prywes
et al, we focus on extension by a single letter. Such an
extension form is sufficient for the digital abstraction
of the emergence of a code as done here in this paper,
and we therefore ignore the option of ligation (which
may accelerate some of the processes described in this
paper, but is not required for their occurrence).
We refer to the process of Single Nucleotide Elon-
gation by Template, as SNET. In particular, when the
SNET is a non enzymatic process, namely, it involves
no enzyme and no ribozyme, we call it NESNET —
Non-Enzymatic Single Nucleotide Elongation by Tem-
plate.
Although the idea of non-motoric extension via
NESNET processes is old and well established, it was
however not very successful; it became a viable direc-
tion only very recently.
For consistence of our model notation with biol-
ogy we identify the letters
r
1
, r
2
, r
3
,
and
r
4
with the
nucleotides
C, A, U, G
, respectively. In RNA,
C
is the
complementary of
G
(and vice versa of course), and
U
and
A
are also complementary to each other. In past
experiments, only the letters
r
1
(
C
) and
r
4
(
G
) were
easily added during a NESNET. In contrast, the letters
r
2
(
A
) and
r
3
(
U
) were not successfully added during
a NESNET. They were added in a very slow rate that
did not allow template replication of functional RNA
sequences faster then they degrade, not even of short
RNA strings containing a very few
U
and
A
letters,
see exact details and several references in (Prywes
et al., 2016). To overcome this problem, researchers
progressing the RNA-world hypothesis often assumed
CG
-only or
CG
-rich RNA-world. Still, no reasonable
NESNET had been performed unless the RNA strings
were fully composed of
C
and
G
. Assume
R
0
is the
complementary of
R
— when a string
R
is already at-
tached to
R
00
which is a part of
R
0
, say from beginning
of
R
till some location, then NESNET, adding the next
letter to
R
00
, can potentially be catalyzed by various
common food molecules: When the next letter in
R
is
C
or
G
, such catalysis for adding a single monomer
(the next letter to
R
00
) is well established. However, to
add U or A seemed nearly impossible, till recently.
The recent finding of (Prywes et al., 2016) is a
new catalytic process in which trimers, length-3
R
-
strings, act as catalysts. More explicitly, (Prywes et al.,
2016) (see also other work done in Szostak’s group)
investigate various catalytic processes enhancing the
probability of attaching the single letter
A
or
U
when
needed. They found that if the
next
trimer (
R
0
trimer
portion of the future
R
00
), right after the letter to be
joint to
R
00
, is attached temporarily by a controlled
supply of
R
0
trimer
, then the probability of adding
A
or
U
is increased by orders of magnitude and became
quite similar to the probability of adding
C
or
G
under
similar conditions. See Figure 2b in (Prywes et al.,
2016).
One drawback of (Prywes et al., 2016) sugges-
tion is that the supply of trimers must be controlled:
Due to competition with other trimers, the next trimer
R
0
trimer
must be present in large quantities relative to
other trimers in order to get attached to the
R
00
string,
and catalyze the NESNET. To facilitate that, (Pry-
wes et al., 2016) used in each of their experiments at
most
4
relevant trimers. They noticed that the pace
of the attachment (and hence the catalysis) is strongly
reduced with the number of different trimers that are
present. Note that the total number of possible trimers
is
64
— to cover all possible combinations of three let-
ters. It thus seems reasonable that the pace of catalysis
became negligible in the natural environment (of the
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
57
prebiotic soup), where all trimers appeared, as food
molecules, in similar quantities.
To overcome this drawback, we suggest briefly
here (and in full details in the journal version of this
work), that the problem can be fully resolved, under
the assumption of a CG-rich world. We observe that
under the limitation that between any two adjacent A
or U nucleotides in an
R
string, there is a sequence
of at least three C or G nucleotides — the number of
required different types of trimers to enable NESNET
catalysis of RNA template replication is decreased
from
64
to
8
. With
8
trimers only it is expected that
the pace will be small but will not be negligible, given
the experimental results with
4
trimers in [See figure 4
of (Prywes et al., 2016)]. Note that in such strings as
described above NESNET by trimers, as in (Prywes
et al., 2016) will only be needed for adding each of the
A
or
U
letters, while
C
or
G
can be added by various
simpler NESNET methods, see references in (Prywes
et al., 2016). Since, in a CG-rich world, most of the
trimers will indeed be made of C and G nucleotides
only — the
8
strings will not have much competition
with the other (rare) strings.
This finding opens the door for template replica-
tion long before a polymerase
P
0
existed, and such a
result is important for almost any RNA-world model,
as well as for COPACS — with or without assuming
RNA-world. In the next section we provide, based
on (Prywes et al., 2016) and on (Agmon and Mor,
2015), a potential step by step evolution from a chemi-
cal era to COPACS.
5 THE EMERGENCE OF A CODE
In this section we describe the main phases of our
offered path from the R and P molecules of the pri-
mordial soup to the emergence of the coded life and
Agmon-Mor COPACS.
5.1 Phases 1, 2 and 3: From Basic
Molecular Evolution to
RNA-Peptides-World (RP-World)
Phase 1
: Assume a world rich of short RNA (
R
)
strings, nucleotides (
r
letters) and amino acids (
p
let-
ters). A longer string can be template replicated via
the non-motoric step, SNET. We may assume a fully
functional RNA-world, where
R
∗
is a ribozyme cat-
alyzing template replication, or we may assume pre-
RNA-world where there are several
R
strings and these
are trimers catalyzing SNET a la Prywes et al.
Phase 2
: Assuming a continuous molecular evolution
(namely step by step), the transformation from phase
1 to phase 2 may be the following. Assume a spe-
cific
R
string acting as a catalyst for binding a single
amino acid to another one or to a short (bi-/tri-) peptide.
Such a string, if emerges, can enhance the environment
by many short peptides. This is speculated to be the
proto-ribosome (
PR
), for example the Agmon-Bashan-
Yonath (ABY) proto-ribosome (Agmon et al., 2006;
Agmon et al., 2009; Agmon, 2009) which we write
here as
PR
-ABY or more generally as
PR
-non-motoric.
The adjective non-motoric is added to clarify that this
PR
— in contrast to today’s
PR
— does not translate
messenger RNA molecules into peptides. It does how-
ever catalyze (thus far — in theory) the creation of
backbone connections between random short peptides
(or a single AA) and an additional AA, and the proba-
bilities for its (the ABY-
PR
) appearance in a prebiotic
world were estimated (Agmon, 2016; Agmon, 2017)
and seem feasible. See Figure 3, where
˜
R
0
is the non
motoric PR.
We emphasize that the ABY proto-ribosome is non
motoric, and furthermore, it does not translate from
messenger RNA to a peptide. One could maybe hope
to define a non-motoric
PR
that does translate from
a messenger RNA to a peptide, but to the best of our
knowledge it is not easy to suggest such a molecule
and it had not yet had been designed or even mentioned
in the literature.
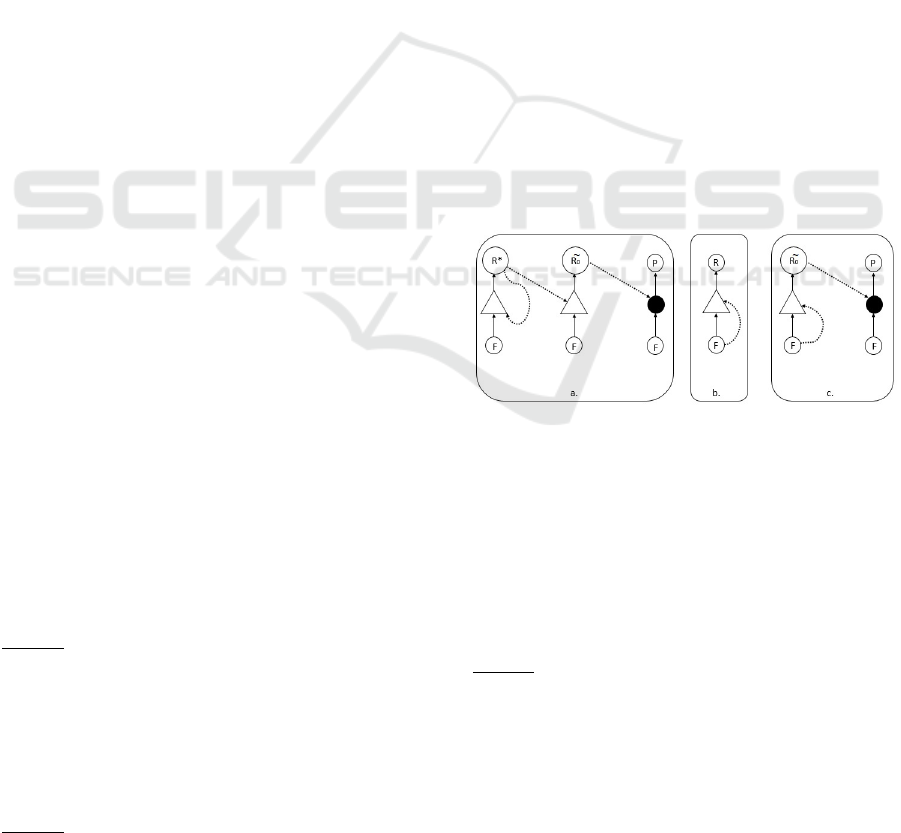
Figure 3: The proto Ribosome (
PR
) appear: a. The proto
ribosome appears and generates many random short peptides.
In this item we assume a ribozyme string
R
∗
that catalyzes
all template replication (including its own). b. Repeating
Figure 2, but this time catalysis is done by short
R
strings
(e.g. trimers) assumed to be part of the food set and not by
some arbitrary
C
molecule. c. Identical to item “a” here,
with the proto ribosome appearance, yet now we assume that
the catalysts for all template replication are several trimers
R
that are hidden in the food set (and not a sophisticated
ribozyme as in item “a”).
Phase 3
: Once the primordial soup is enriched by
many short
P
strings, various
R
strings potentially
evolve and create constructive interactions with pep-
tides attached to them, for example by improving catal-
ysis. Since the added
PR
indirectly enhance various
catalytic processes, various ACS may be formed. We
do not attempt to specify such ACS here, as most of the
generated molecules are not directly relevant for the
COMPLEXIS 2018 - 3rd International Conference on Complexity, Future Information Systems and Risk
58
next steps, and as such ACS might not be sufficiently
stable because
˜
R
0
generates random peptides.
Now, assume that one specific peptide
P
∗
generated
by the
PR
enhances the template replication catalysis
done by the catalyst
R
∗
or the SNET catalyzed by the
relevant set of trimers
R
(being food molecules), see
Figure 4.
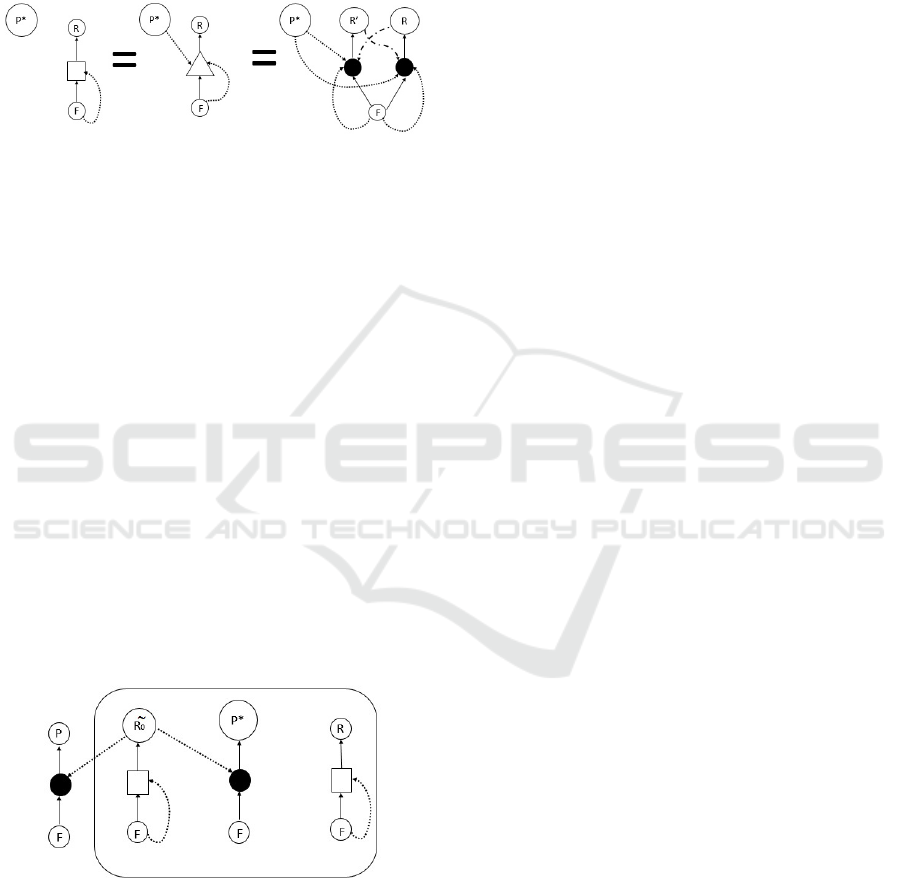
Figure 4: Defining the square-process notation — when
P
∗
exists: The square notation is defined here to replace the tri-
angle whenever catalysis of template replication is enhanced
by the catalyst
P
∗
. Note that the use of two catalysts (as is
explicit in the middle and in the right side of the equation)
means that catalysis is much stronger with both, and might
exist (yet be weaker) or not exist at all if just one of the two
catalysts is in the set.We do not add the catalysis line from
P
∗
in the left figure and in the later figures, because
P
∗
is a
catalyst for all
R
strings, and adding many lines just makes
the figures unclear and cumbersome. Instead we upgrade the
triangle notation to square notation such that each square is
catalyzed by P
∗
without this being explicitly denoted.
Now the
PR
plus the
P
∗
form an ACS, see Figure 5.
Note however that this ACS is still not a stable one, be-
cause the
PR
generates many other peptides (e.g.
P
in
Figure 5) hence wasting or even exhausting the
p
food,
and the
P
∗
helps template replication of all strings, not
just the
PR
(e.g.
R
in Figure 5), hence exhausting the
r
food. We now reached phase 3 of our offered path of
the molecular evolution, in the direction of generating
our COPACS. The dual catalysis by trimers plus
P
∗
is
assumed much stronger than a single catalysis by just
one of those.
Figure 5: If
P
∗
is generated by the proto ribosome we obtain
a (relatively unstable) ACS:
r
letters (and trimers) food is
wasted on
P
∗
catalyzing random
R
strings, and
p
letters food
is wasted on the proto ribosome catalyzing random short
P
strings.
Here are some major options for how the peptide
P
∗
helps the template replication process, the first two
options are the most promising in our eyes:
1.
Probably the most promising direction could be
helping the template
R
and its replication
R
0
to split
from each other, or in other words — preventing
the re-joining (Rouch, 2014; Jia et al., 2016) of
the two strings when they start to split up. It may
well be that the strings will not split at all, or that it
will take an extremely long time for them to split,
unless the split is supported. And
P
∗
could be a
short peptide supporting this split, a split that may
start at one end, while the SNET is still applied to
approach the other end.
2.
The second most promising direction in our eyes
could be that
P
∗
directly replaces the trimers in per-
forming SNET. This suggestion partially recovers
the original COPACS idea of (Agmon and Mor,
2015), yet with
P
∗
being a non-motoric peptide
that helps a single non-motoric extension at a time.
Namely, after each step the peptide
P
∗
leaves, and
then a different one or the same one comes back
for the next step. This is in contrast to the sin-
gle motoric polymerase called
P
0
in (Agmon and
Mor, 2015). We may still call
P
∗
in this case a
polymerase, however in this case — a non-motoric
polymerase, of course.
3.
A bit similar to the first option above, yet, probably
a much less promising direction could be helping
the template
R
and its replication
R
0
not to split
too fast. If the splitting is too fast, the replication
might end while only a part of R
0
is formed.
4.
The peptide
P
∗
might help trimers that needs to
leave after catalyzing the SNET to leave much
faster, or might help irrelevant trimers that are not
suppose to get attached there, to leave much faster
and free the space for the relevant trimers. Alterna-
tively, the peptide might help the relevant trimers
to attach to the template or to become activated by
bonding to the correct molecule; see (Prywes et al.,
2016) for a discussion of the activated trimers via
different activating molecules bonded to them.
5.
We may assume
P
∗
directly replaces the trimers in
catalyzing template replication, and moving one
step at a time. This suggestion fully recovers the
original COPACS idea of (Agmon and Mor, 2015);
its disadvantage is that probably the required mo-
toric peptide (named proto-polymerase in (Agmon
and Mor, 2015)) in this case is supposed to be
much longer than any other option for
P
∗
above,
hence assuming a very long and quite specific mes-
senger RNA (even if we consider many options
for it, allowing the maximal possible flexibility
in the monomers choice of both the peptide and
its corresponding RNA) and the probability to its
spontaneous emergence is low — see phase 5.
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
59
The later figures in the paper describe a scenario in
which both
P
∗
and the trimers are involved in the
catalysis. Only the second and the last scenario devi-
ate from this since in these options
P
∗
fully replaces
the trimers [and hence the discussion in these options
just merges with the discussion in (Agmon and Mor,
2015)].
Note that we only gave a few examples of what
the first code could have been. Much more research
is needed in order to support one option over another,
and for sure many other options can be offered, for
short peptides that would help the SNET.
5.2 Phases 4 and 5: From
RNA-Peptides-World to COPACS
Phase 4
: The non motoric
PR
(seen as
˜
R
0
in previous
figures) does not lead to a stable ACS. Three more
steps are required in order to yield a stable ACS that
should contain a motoric proto ribosome (
R
0
), and
have stability: First, the
p
letters joint by the non mo-
toric
˜
R
0
need to be upgraded to have short RNA legs
attached to them. A molecule synthesizing bonding be-
tween a single amino acid and a short RNA string can
be named proto-synthetase, and it is commonly agreed
that various short molecules (short RNA strings, very
short peptides, etc) may synthesize such a bond in a
non-selective way, even if not very efficiently (Schim-
mel and de Pouplana, 1995).
Let us add to that picture also a stereochemical
attraction between specific amino acids and specific
RNA-triplets (named codons or anti-codons, for later
use in phase 5). If we assume that sometimes the
attached RNA strings might be RNA-helix (in their
shape), we get proto-tRNA charged with amino acids.
The molecule responsible for synthesizing that charg-
ing may be a food molecule or may be, as in Figure 6,
a short ribozyme (not in the food set).
Phase 5
: This is the most important and mysterious
step, although it seems to be a vital step in any model
for the origin of life on Earth: At some point the non
motoric PR had to become motoric!
For simplicity, one may assume that first the non-
motoric
PR
(e.g. the ABY-
PR
) already existed, and
then another RNA molecule got attached to it to form a
motoric
PR
. It seems that the ABY-
PR
is contained in
today’s LSU (the Large Sub Unit of the ribosome), and
that today’s SSU (the Small Sub Unit of the ribosome)
had been attached later in evolution. It also seems
that today’s SSU and today’s LSU, together, take care
of the motorics of the current ribosome. We are not
aware of research work explaining the motorics of the
proto-ribosome during the origins of translation.
The motoric
PR
, denoted as
R
0
in (Agmon and
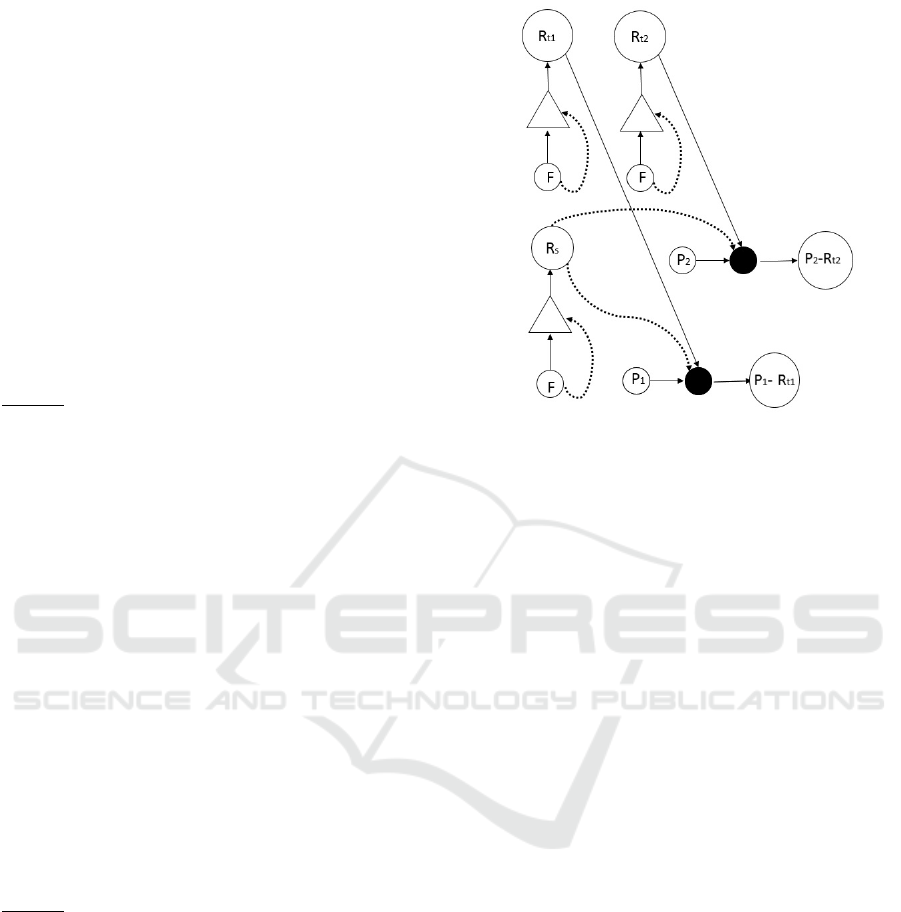
Figure 6: Appearance of
tRNA
strings: This figure presents
the appearance of proto-
tRNA
strings, of charged proto-
tRNA
strings and of the food or non-food molecule syn-
thesizing the charging (non-food
R
string named
R
s
, in this
figure). We only show two
tRNA
s yet there are at least four
in the origin of life.
Mor, 2015) and here, plays a unique role: If an
R
string
(named in this case —
R
mes
) passes through it, every
triplet of r letters (named a coding triplet in this case)
in
R
mes
moves somehow through a “reading” position
in it, probably with the help of
tRNA
attached to such
a coding triple. When the triplet is in the reading
position, the relevant
p
letter attached to the tRNA
tail gets near another
p
letter (or already a formed
short peptide) attached as well to a nearby
tRNA
held
in a nearby location in
R
0
, while still probably also
attached to the
R
mes
. Then the
˜
R
0
part of the PR (see
Phase 2) attaches the
p
letter to the one earlier arrived
(or the already formed short peptide). It makes sense
that a triplet in the tail is attached if it is complementary
to a triplet in R
mes
, hence it is more efficient to attract
charged
tRNA
than to attract directly an amino acid or
a non-charged tRNA.
Note that the
R
string
R
mes
is assumed (for now)
to be random. Each of its triplets is thus translated
to an amino acid, and the entire
R
mes
string is trans-
lated to a specific corresponding short peptide. The
P
string built by this “translation” process is random in
sequence, but is uniquely dictated by the string
R
mes
(three letters after three). In some sense, the string
R
mes
acts as a template for building a specific string
P
,
hence we name it
R
mes
(P)
. Let us refer to this type of
templating operation as “translation”, the term used for
this process in biology, and still denote it by a template
line in the relevant figures.
In most cases the strings
R
mes
(P)
and hence also
COMPLEXIS 2018 - 3rd International Conference on Complexity, Future Information Systems and Risk
60
the resulting
P
strings are not useful, although they
could enrich the local environment. Suppose that just
ONCE
, a string
R
mes
(P
∗
)
appears and go through the
motoric proto ribosome. Namely,
P
∗
of Phase 3 is then
generated via translation. Once this occurs, the set of
three strings (together with charged tRNAs and the
synthetase), does the following: The string
R
mes
(P
∗
)
is template duplicated using the help from
P
∗
, hence
more
R
mes
(P
∗
)
will appear. The strings building
R
0
are
also template duplicated using the help from
P
∗
, hence
more
R
0
will appear. More such strings
R
mes
(P
∗
)
will
move via the generated (motoric)
R
0
hence more
P
∗
will also appear.
This scenario now leads to the emergence of a code:
R
mes
(P
∗
)
is the code-word that contains the informa-
tion concerned with the sequence of the
p
letters in the
P
∗
string. The set
P
∗
,
R
0
, and this unique
R
mes
(P
∗
)
,
is an ACS: to be more precise, it leads then to the
generation of the complementary strings of
R
0
and of
R
mes
(P
∗
)
(as the first two
R
strings are expected to be
in the vicinity of
P
∗
), and this addition, along with the
tRNA
and synthetase (that are short and hence already
highly common in the environment) completes a code
prompting ACS — COPACS; see Figure 7.
Once such a COPACS is built, it becomes more and
more prevalent, inside the compartment (this is true
for any ACS that does not include a suicidal catalyst
(Vasas et al., 2012)), if sufficient food molecules are
available (in contrast to the case of non-stable ACS
due to the food been used by many other molecules
and hence exhausted).
By diffusion (Chen and Nowak, 2012), and the de-
struction and construction of compartment walls, the
environment (including also neighboring and newly
formed compartments) can be potentially enriched
with these COPACS strings.
5.3 Phase 6: From Our COPACS to
Agmon-Mor COPACS
Far later in the evolution, once
R
0
is common, and
various
R
mes
encode various peptides, there evolve
a set of two special strings. A unique
R
mes
string,
that encodes
P
0
is added (just once), and it generates
a unique string called the polymerase,
P
0
, or more
correctly, the proto-polymerase (Lazcano et al., 1988;
Aravind et al., 2002; Iyer et al., 2003). This proto-
polymerase catalyzes the template replication of any
R
string that gets close to it. Once this occurs, the set
of three strings
P
0
,
R
0
and
R
mes
(P
0
)
along with tRNAs,
charged tRNAs and the synthetase, forms the COPACS
suggested by Agmon-Mor (Agmon and Mor, 2015).
For the transformation to selective synthetase and
the fixation of the code see (Agmon and Mor, 2015).
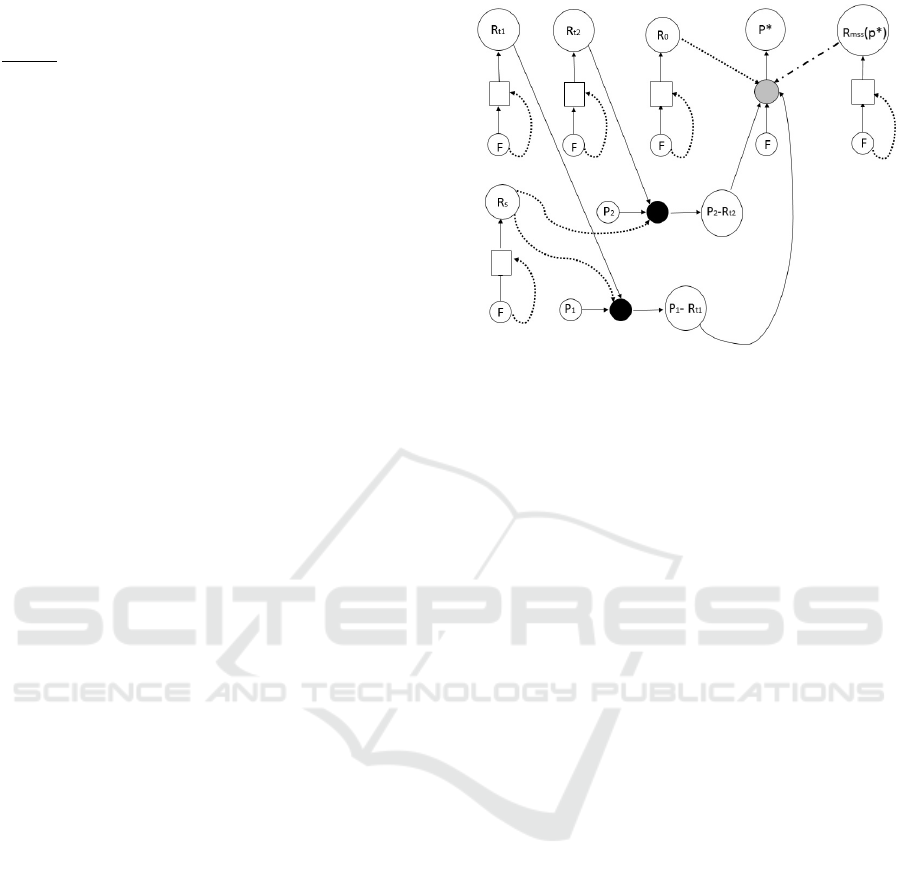
Figure 7: Appearance of the COPACS: This figure presents
the appearance of our COPACS, in which the players are
P
∗
and its coding
R
mes
, the motoric
PR
—
R
0
, the strings
tRNA
s and charged
tRNA
s, the synthetase (being a non-food
molecule here), and the trimers
R
∗
responsible (along with
P
∗
) for template replication. Note that the translation is also
shown via a template line, as there is one to one correspon-
dence between the coding triplets in
R
mes
and the resulting
peptide. Note also a new notation, the gray circle, to denote
that not necessarily all charged
tRNA
s must contribute to
building a peptide.
6 DISCUSSION
In this paper we aimed, by abstracting components
from current biology, to put forward a feasible model
(relying on a continuous evolution) for the emergence
of life as we know it, that is - life rooted in the genetic
code. In this model we show how RNA molecules and
amino-acids form polymers that create Code Prompt-
ing Auto Catalytic Set of molecules - COPACS. CO-
PACS were first suggested in AM15. However, their
COPACS seemed to rely on the joint appearance of
two relatively complex R strings, a motoric ribosome,
and a messenger RNA encoding a motoric peptide,
the polymerase. Such a joint event seems to be rather
unlikely.
The COPACS suggested here were derived by tak-
ing into account a novel method suggested and ex-
plored by Prywes et al, a method of template replica-
tion via non-motoric catalysis causing SNET. Based
on that possibility, in which the catalyst of the template
replication does not need to be motoric anymore, we
suggest a much more realistic COPACS. We provide
several alternatives for the emergence of a vital protein
component in an ACS hence suggesting simple sug-
gestions for the first code word in a COPACS, without
relying on the emergence of a complex motoric poly-
merase. The scenario we presented here, although of
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
61

course still speculative, clarifies that continuous evolu-
tion of ACS could lead to the emergence of the genetic
code.
Our COPACS hypothesis does not contradict the
prior existence of an “RNA world” (Woese, 1967;
Crick, 1968; Orgel, 1968; Gilbert, 1986). In this
widely accepted hypothesis concerned with the ori-
gin of life, a “world” where RNA enzymes acted as
the sole catalysts preceded life as we know it (where
the majority of catalysis is performed by proteins). An
RNA-world would have required a replicase built of
RNA that could have copied itself as well as the other
functional ribozymes, together forming a non-coded
ACS. The method of Prywes et al allows closing a
serious gap in the RNA-world hypothesis, by avoiding
the need for a motoric RNA-based replicase. The CO-
PACS in our model could have emerged and started
functioning within an RNA-world, providing a possi-
ble missing link between the RNA-world and an RNA-
protein world, which required a transformation, from
replication by an RNA enzyme to (RNA) replication by
a protein enzyme. Alternatively, such COPACS could
have materialized spontaneously without the phase
of RNA world, i.e. before any complex replicative
molecular system existed except simple SNET and/or
ligation of a few r letters at a time.
We expect future research to further investigate
the main players of our model: to improve knowl-
edge regarding non-motoric SNET, to prove that some
peptides enhance this SNET. Another major goal that
would make the model much more relevant in current
lab experiments may be to investigate the possibility
of a non-motoric
PR
that still can perform translation,
similarly to how
R
trimers and
P
∗
perform the non-
motoric SNET, by arriving to a site and leaving it.
ACKNOWLEDGMENTS
We thank the Israeli Ministry of Defense Research and
Technology Unit. We thank Yoram Gerchman and
Yuval Elias for interesting discussions and comments.
We especially thank Ilana Agmon for numerous dis-
cussions, comments and insights.
REFERENCES
Agmon, I. (2009). The dimeric proto-ribosome: Structural
details and possible implications on the origin of life.
Int. J. Mol. Sci., 10:2921–2934.
Agmon, I. (2016). Could a proto-ribosome emerge sponta-
neously in the prebiotic world? Molecules, 21:1701.
Agmon, I. (2017). Sequence complementarity at the riboso-
mal peptidyl transferase centre implies self-replicating
origin. FEBS Letters.
Agmon, I., Bashan, A., and Yonath, A. (2006). On ribosome
conservation and evolution. Israel Journal of Ecology
and Evolution, 52:359–374.
Agmon, I., Davidovich, C., Bashan, A., and Yonath, A.
(2009). Identification of the prebiotic translation
apparatus within the contemporary ribosome. See
http://precedings.nature.com/documents/2921/version/1.
Agmon, I. and Mor, T. (2015). A model for the emergence
of coded life. TPNC 2015, LNCS, 9477:97–108.
Aravind, L., Mazumder, R., Vasudevan, S., and Koonin,
E. V. (2002). Trends in protein evolution inferred from
sequence and structure analysis. Curr. Opin. Struct.
Biol., 12:392–399.
Chen, I. A. and Nowak, M. A. (2012). From prelife to life:
How chemical kinetics become evolutionary dynamics.
Acc. Chem. Res., 45:2088–2096.
Crick, F. H. C. (1968). The origin of the genetic code. J.
Molec. Biol., 38:367–379.
Dyson, F. J. (1985). Origins of life. Cambridge University
Press.
Ferris, J. P. (2002). Montmorillonite catalysis of 30–50 mer
oligonucleotides: laboratory demonstration of potential
steps in the origin of the RNA world. Orig. Life Evol.
Biosph., 32:311–332.
Gilbert, W. (1986). Origin of life: The RNA world. Nature,
319:618.
Guerrier-Takada, C., Gardiner, K., Marsh, T., Pace, N., and
Altman, S. (1983). The RNA moiety of ribonuclease
p is the catalytic subunit of the enzyme. Cell, 35:849–
857.
Hordijk, W., Hein, J., and Steel, M. (2010). Autocatalytic
sets and the origin of life. Entropy, 12:1733–1742.
Hordijk, W., Kauffman, S. A., and Steel, M. (2011). Re-
quired levels of catalysis for emergence of autocatalytic
sets in models of chemical reaction systems. Int. J. Mol.
Sci., 12:3085–3101.
Hordijk, W. and Steel, M. (2004). Detecting autocatalytic,
self-sustaining sets in chemical reaction systems. J.
Theor. Biol., 227:451–461.
Horning, D. P. and Joyce, G. F. (2016). Amplification of
RNA by an RNA polymerase ribozyme. Proc. Natl.
Acad. Sci. USA, 113:9786–9791.
Ikehara, K. (2005). Possible steps to the emergence of life:
The [GADV]-protein world hypothesis. Chem. Rec.,
5:107–118.
Iyer, L. M., Koonin, E. V., and Aravind, L. (2003). Evo-
lutionary connection between the catalytic subunits
of DNA-dependent RNA polymerases and eukaryotic
RNA-dependent RNA polymerases and the origin of
RNA polymerases. BMC Struct. Biol., 3:1–23.
Jia, T. Z., Fahrenbach, A. C., Kamat, N. P., Adamala, K. P.,
and Szostak, J. W. (2016). Oligoarginine peptides
slow strand annealing and assist non-enzymatic RNA
replication. NC, 8:915–921.
Johnson, D. B. and Wang, L. (2010). Imprints of the genetic
code in the ribosome. Proc. Natl. Acad. Sci. USA,
107:8298–8303.
COMPLEXIS 2018 - 3rd International Conference on Complexity, Future Information Systems and Risk
62

Kauffman, S. (1986). Autocatalytic sets of proteins. J. Theor.
Biol., 119:1–24.
Kauffman, S. (2011). Approaches to the origin of life on
earth. Life, 1:34–48.
Koonin, E. V. and Martin, W. (2005). On the origin of
genomes and cells within inorganic compartments.
TRENDS Genet., 21:647–654.
Koonin, E. V. and Novozhilov, A. S. (2009). Origin and
evolution of the genetic code: the universal enigma.
IUBMB Life, 61:99–111.
Koshland, D. E. (2002). The seven pillars of life. Science,
295:2215–2216.
Kunin, V. (2000). A system of two polymerases – a model for
the origin of life. Orig. Life Evol. Biosph., 30:459–466.
Lahav, N., Nir, S., and Elitzur, A. (2001). The emergence of
life on earth. Prog. Biol. Molec. Biol., 75:75–120.
Lane, N. (2015). The vital question: energy, evolution, and
the origins of complex life. W.W. Norton & Company,
N.Y., USA.
Lane, N. and Martin, W. (2012). The origin of membrane
bioenergetics. Cell, 151:1406–1416.
Lazcano, A., Fastag, J., Gariglio, P., Ram
´
ırez, C., and Or
´
o, J.
(1988). On the early evolution of RNA polymerase. J.
Molec. Evol., 27:365–376.
Mast, C. B. and Braun, D. (2010). Thermal trap for DNA
replication. Phys. Rev. Lett., 104:188102.
Orgel, L. E. (1968). Evolution of the genetic apparatus. J.
Molec. Biol., 38:381–393.
Prywes, N., Blain, J., Del Frate, F., and Szostak, J. (2016).
Nonenzymatic copying of RNA templates contain-
ing all four letters is catalyzed by activated oligonu-
cleotides. eLife, 5.
Rouch, A. (2014). Evolution of the first genetic cells and
the universal genetic code: A hypothesis based on
macromolecular coevolution of RNA and proteins. J.
Theor. Biol., 357:220–244.
Schimmel, P. and de Pouplana, L. R. (1995). Transfer RNA:
from minihelix to genetic code. Cell, 81:983–986.
Schroedinger, E. (1944). What is life? The physical aspect
of the living cell. Cambridge University Press.
Segr
´
e, D., Ben-Eli, D., Deamer, D. W., and Lancet, D. (2001).
The lipid world. Orig. Life Evol. Biosph., 31:119–145.
van der Gulik, P., Massar, S., Gilis, D., Buhrman, H., and
Rooman, M. (2009). The first peptides: the evolution-
ary transition between prebiotic amino acids and early
proteins. J. Theor. Biol., 261:531–539.
Vasas, V., Fernando, C., Santos, M., Kauffman, S., and
Szathm
´
ary, E. (2012). Evolution before genes. Biol.
Direct, 7:1–14.
Watson, J. D. and Crick, F. H. (1953). Genetical implica-
tions of the structure of deoxyribonucleic acid. Nature,
171:964–967.
Woese, C. R. (1965). On the evolution of the genetic code.
Proc. Natl. Acad. Sci. USA, 54:1546.
Woese, C. R. (1967). The genetic code: the molecular basis
for genetic expression. Harper and Row, New York.
Yarus, M., Caporaso, J. G., and Knight, R. (2005). Origins
of the genetic code: the escaped triplet theory. Annu.
Rev. Biochem., 74:179–198.
On Code-Prompting Auto-Catalytic Sets and the Origins of Coded Life
63
